Prokaryotic chaperones support yeast prions and thermotolerance and define disaggregation machinery interactions
- PMID: 22732191
- PMCID: PMC3430535
- DOI: 10.1534/genetics.112.142307
Prokaryotic chaperones support yeast prions and thermotolerance and define disaggregation machinery interactions
Abstract
Saccharomyces cerevisiae Hsp104 and Escherichia coli ClpB are Hsp100 family AAA+ chaperones that provide stress tolerance by cooperating with Hsp70 and Hsp40 to solubilize aggregated protein. Hsp104 also remodels amyloid in vitro and promotes propagation of amyloid prions in yeast, but ClpB does neither, leading to a view that Hsp104 evolved these activities. Although biochemical analyses identified disaggregation machinery components required for resolubilizing proteins, interactions among these components required for in vivo functions are not clearly defined. We express prokaryotic chaperones in yeast to address these issues and find ClpB supports both prion propagation and thermotolerance in yeast if it is modified to interact with yeast Hsp70 or if E. coli Hsp70 and its cognate nucleotide exchange factor (NEF) are present. Our findings show prion propagation and thermotolerance in yeast minimally require cooperation of species-specific Hsp100, Hsp70, and NEF with yeast Hsp40. The functions of this machinery in prion propagation were directed primarily by Hsp40 Sis1p, while thermotolerance relied mainly on Hsp40 Ydj1p. Our results define cooperative interactions among these components that are specific or interchangeable across life kingdoms and imply Hsp100 family disaggregases possess intrinsic amyloid remodeling activity.
Figures




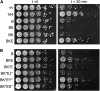
Similar articles
-
Hsp70 targets Hsp100 chaperones to substrates for protein disaggregation and prion fragmentation.J Cell Biol. 2012 Aug 6;198(3):387-404. doi: 10.1083/jcb.201201074. J Cell Biol. 2012. PMID: 22869599 Free PMC article.
-
Chaperone networks in protein disaggregation and prion propagation.J Struct Biol. 2012 Aug;179(2):152-60. doi: 10.1016/j.jsb.2012.05.002. Epub 2012 May 10. J Struct Biol. 2012. PMID: 22580344 Review.
-
Hsp40s specify functions of Hsp104 and Hsp90 protein chaperone machines.PLoS Genet. 2014 Oct 16;10(10):e1004720. doi: 10.1371/journal.pgen.1004720. eCollection 2014 Oct. PLoS Genet. 2014. PMID: 25329162 Free PMC article.
-
Bacterial and Yeast AAA+ Disaggregases ClpB and Hsp104 Operate through Conserved Mechanism Involving Cooperation with Hsp70.J Mol Biol. 2016 Oct 23;428(21):4378-4391. doi: 10.1016/j.jmb.2016.09.003. Epub 2016 Sep 9. J Mol Biol. 2016. PMID: 27616763
-
Towards a unifying mechanism for ClpB/Hsp104-mediated protein disaggregation and prion propagation.Biochem Cell Biol. 2010 Feb;88(1):63-75. doi: 10.1139/o09-118. Biochem Cell Biol. 2010. PMID: 20130680 Review.
Cited by
-
How Do Yeast Cells Contend with Prions?Int J Mol Sci. 2020 Jul 3;21(13):4742. doi: 10.3390/ijms21134742. Int J Mol Sci. 2020. PMID: 32635197 Free PMC article. Review.
-
Antiprion systems in yeast cooperate to cure or prevent the generation of nearly all [PSI+] and [URE3] prions.Proc Natl Acad Sci U S A. 2022 Jul 12;119(28):e2205500119. doi: 10.1073/pnas.2205500119. Epub 2022 Jul 5. Proc Natl Acad Sci U S A. 2022. PMID: 35787049 Free PMC article.
-
Mutations Outside the Ure2 Amyloid-Forming Region Disrupt [URE3] Prion Propagation and Alter Interactions with Protein Quality Control Factors.Mol Cell Biol. 2020 Oct 13;40(21):e00294-20. doi: 10.1128/MCB.00294-20. Print 2020 Oct 13. Mol Cell Biol. 2020. PMID: 32868289 Free PMC article.
-
Heat shock protein 104 (Hsp104)-mediated curing of [PSI+] yeast prions depends on both [PSI+] conformation and the properties of the Hsp104 homologs.J Biol Chem. 2017 May 26;292(21):8630-8641. doi: 10.1074/jbc.M116.770719. Epub 2017 Apr 3. J Biol Chem. 2017. PMID: 28373280 Free PMC article.
-
Stand-alone ClpG disaggregase confers superior heat tolerance to bacteria.Proc Natl Acad Sci U S A. 2018 Jan 9;115(2):E273-E282. doi: 10.1073/pnas.1712051115. Epub 2017 Dec 20. Proc Natl Acad Sci U S A. 2018. PMID: 29263094 Free PMC article.
References
-
- Acebron S. P., Fernandez-Saiz V., Taneva S. G., Moro F., Muga A., 2008. DnaJ recruits DnaK to protein aggregates. J. Biol. Chem. 283: 1381–1390 - PubMed
-
- Chernoff Y. O., Lindquist S. L., Ono B., Inge-Vechtomov S. G., Liebman S. W., 1995. Role of the chaperone protein Hsp104 in propagation of the yeast prion-like factor [psi+] Science 268: 880–884 - PubMed
-
- Cox B. S., 1965. “PSI” a cytoplasmic suppressor of super-suppressor in yeast. Heredity 20: 505–521
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

