Endo-exo synergism in cellulose hydrolysis revisited
- PMID: 22733813
- PMCID: PMC3436550
- DOI: 10.1074/jbc.M112.381624
Endo-exo synergism in cellulose hydrolysis revisited
Abstract
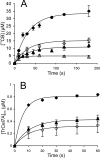
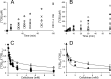
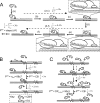
Synergistic cooperation of different enzymes is a prerequisite for efficient degradation of cellulose. The conventional mechanistic interpretation of the synergism between randomly acting endoglucanases (EGs) and chain end-specific processive cellobiohydrolases (CBHs) is that EG-generated new chain ends on cellulose surface serve as starting points for CBHs. Here we studied the hydrolysis of bacterial cellulose (BC) by CBH TrCel7A and EG TrCel5A from Trichoderma reesei under both single-turnover and "steady state" conditions. Unaccountable by conventional interpretation, the presence of EG increased the rate constant of TrCel7A-catalyzed hydrolysis of BC in steady state. At optimal enzyme/substrate ratios, the "steady state" rate of synergistic hydrolysis became limited by the velocity of processive movement of TrCel7A on BC. A processivity value of 66 ± 7 cellobiose units measured for TrCel7A on (14)C-labeled BC was close to the leveling off degree of polymerization of BC, suggesting that TrCel7A cannot pass through the amorphous regions on BC and stalls. We propose a mechanism of endo-exo synergism whereby the degradation of amorphous regions by EG avoids the stalling of TrCel7A and leads to its accelerated recruitment. Hydrolysis of pretreated wheat straw suggested that this mechanism of synergism is operative also in the degradation of lignocellulose. Although both mechanisms of synergism are used in parallel, the contribution of conventional mechanism is significant only at high enzyme/substrate ratios.
Figures



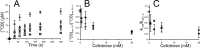
References
-
- Ragauskas A. J., Williams C. K., Davison B. H., Britovsek G., Cairney J., Eckert C. A., Frederick W. J., Jr., Hallett J. P., Leak D. J., Liotta C. L., Mielenz J. R., Murphy R., Templer R., Tschaplinski T. (2006) The path forward for biofuels and biomaterials. Science 311, 484–489 - PubMed
-
- Himmel M. E., Ding S. Y., Johnson D. K., Adney W. S., Nimlos M. R., Brady J. W., Foust T. D. (2007) Biomass recalcitrance: engineering plants and enzymes for biofuels production. Science 315, 804–807 - PubMed
-
- Chundawat S. P., Beckham G. T., Himmel M. E., Dale B. E. (2011) Deconstruction of lignocellulosic biomass to fuels and chemicals. Annu. Rev. Chem. Biomol. Eng. 2, 121–145 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

