Trim71 cooperates with microRNAs to repress Cdkn1a expression and promote embryonic stem cell proliferation
- PMID: 22735451
- PMCID: PMC3518406
- DOI: 10.1038/ncomms1909
Trim71 cooperates with microRNAs to repress Cdkn1a expression and promote embryonic stem cell proliferation
Abstract
Pluripotent embryonic stem cells have a shortened cell cycle that enables their rapid proliferation. The embryonic stem cell-specific miR-290 and miR-302 microRNA families promote proliferation whereas let-7 microRNAs inhibit self-renewal, and promote cell differentiation. Lin28 suppresses let-7 expression in embryonic stem cells. Here to gain further insight into mechanisms controlling embryonic stem cell self-renewal, we explore the molecular and cellular role of the let-7 target Trim71 (mLin41). We show that Trim71 associates with Argonaute2 and microRNAs, and represses expression of Cdkn1a, a cyclin-dependent kinase inhibitor that negatively regulates the G1-S transition. We identify protein domains required for Trim71 association with Argonaute2, localization to P-bodies, and for repression of reporter messenger RNAs. Trim71 knockdown prolongs the G1 phase of the cell cycle and slows embryonic stem cell proliferation, a phenotype that was rescued by depletion of Cdkn1a. Thus, we demonstrate that Trim71 is a factor that facilitates the G1-S transition to promote rapid embryonic stem cell self-renewal.
Figures
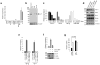

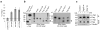




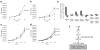
References
-
- Winter J, et al. Many roads to maturity: microRNA biogenesis pathways and their regulation. Nat Cell Biol. 2009;11(3):228–234. - PubMed
-
- Hammond SM, et al. Argonaute2, a link between genetic and biochemical analyses of RNAi. Science. 2001;293(5532):1146–1150. - PubMed
-
- Liu J, et al. Argonaute2 is the catalytic engine of mammalian RNAi. Science. 2004;305(5689):1437–1441. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

