Equilibrium unfolding of the PDZ domain of β2-syntrophin
- PMID: 22735534
- PMCID: PMC3379018
- DOI: 10.1016/j.bpj.2012.05.021
Equilibrium unfolding of the PDZ domain of β2-syntrophin
Abstract
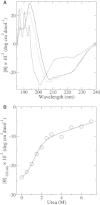
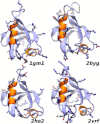
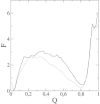
β2-syntrophin, a dystrophin-associated protein, plays a pivotal role in insulin secretion by pancreatic β-cells. It contains a PDZ domain (β2S-PDZ) that, in complex with protein-tyrosine phosphatase ICA512, anchors the dense insulin granules to actin filaments. The phosphorylation state of β2-syntrophin allosterically regulates the affinity of β2S-PDZ for ICA512, and the disruption of the complex triggers the mobilization of the insulin granule stores. Here, we investigate the thermal unfolding of β2S-PDZ at different pH and urea concentrations. Our results indicate that, unlike other PDZ domains, β2S-PDZ is marginally stable. Thermal denaturation experiments show broad transitions and cold denaturation, and a two-state model fit reveals a significant unfolded fraction under physiological conditions. Furthermore, T(m) and T(max) denaturant-dependent shifts and noncoincidence of melting curves monitored at different wavelengths suggest that two-state and three-state models fail to explain the equilibrium data properly and are in better agreement with a downhill scenario. Its higher stability at pH >9 and the results of molecular dynamics simulations indicate that this behavior of β2S-PDZ might be related to its charge distribution. All together, our results suggest a link between the conformational plasticity of the native ensemble of this PDZ domain and the regulation of insulin secretion.
Copyright © 2012 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures





Similar articles
-
β2-Syntrophin is a Cdk5 substrate that restrains the motility of insulin secretory granules.PLoS One. 2010 Sep 23;5(9):e12929. doi: 10.1371/journal.pone.0012929. PLoS One. 2010. PMID: 20886068 Free PMC article.
-
The receptor tyrosine phosphatase-like protein ICA512 binds the PDZ domains of beta2-syntrophin and nNOS in pancreatic beta-cells.Eur J Cell Biol. 2000 Sep;79(9):621-30. doi: 10.1078/0171-9335-00095. Eur J Cell Biol. 2000. PMID: 11043403
-
Dephosphorylation of beta2-syntrophin and Ca2+/mu-calpain-mediated cleavage of ICA512 upon stimulation of insulin secretion.EMBO J. 2001 Aug 1;20(15):4013-23. doi: 10.1093/emboj/20.15.4013. EMBO J. 2001. PMID: 11483505 Free PMC article.
-
Interaction of muscle and brain sodium channels with multiple members of the syntrophin family of dystrophin-associated proteins.J Neurosci. 1998 Jan 1;18(1):128-37. doi: 10.1523/JNEUROSCI.18-01-00128.1998. J Neurosci. 1998. PMID: 9412493 Free PMC article.
-
Syntrophins entangled in cytoskeletal meshwork: Helping to hold it all together.Cell Prolif. 2019 Mar;52(2):e12562. doi: 10.1111/cpr.12562. Epub 2018 Dec 4. Cell Prolif. 2019. PMID: 30515904 Free PMC article. Review.
Cited by
-
Single-Molecule Protein Interactions and Unfolding Revealed by Plasmon-Enhanced Fluorescence.Anal Chem. 2025 Jul 29;97(29):15651-15657. doi: 10.1021/acs.analchem.5c01091. Epub 2025 Jul 19. Anal Chem. 2025. PMID: 40682813 Free PMC article.
-
Syntrophin proteins as Santa Claus: role(s) in cell signal transduction.Cell Mol Life Sci. 2013 Jul;70(14):2533-54. doi: 10.1007/s00018-012-1233-9. Epub 2012 Dec 21. Cell Mol Life Sci. 2013. PMID: 23263165 Free PMC article. Review.
-
The order of PDZ3 and TrpCage in fusion chimeras determines their properties-a biophysical characterization.Protein Sci. 2021 Aug;30(8):1653-1666. doi: 10.1002/pro.4107. Epub 2021 Jun 3. Protein Sci. 2021. PMID: 33969912 Free PMC article.
-
Phosphorylation-induced changes in the PDZ domain of Dishevelled 3.Sci Rep. 2021 Jan 15;11(1):1484. doi: 10.1038/s41598-020-79398-5. Sci Rep. 2021. PMID: 33452274 Free PMC article.
References
-
- Adams M.E., Dwyer T.M., Froehner S.C. Mouse α1- and β2-syntrophin gene structure, chromosome localization, and homology with a discs large domain. J. Biol. Chem. 1995;270:25859–25865. - PubMed
-
- Ort T., Maksimova E., Solimena M. The receptor tyrosine phosphatase-like protein ICA512 binds the PDZ domains of β2-syntrophin and nNOS in pancreatic β-cells. Eur. J. Cell Biol. 2000;79:621–630. - PubMed
-
- Lumeng C., Phelps S., Chamberlain J.S. Interactions between β2-syntrophin and a family of microtubule-associated serine/threonine kinases. Nat. Neurosci. 1999;2:611–617. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

