A high-throughput next-generation sequencing-based method for detecting the mutational fingerprint of carcinogens
- PMID: 22735701
- PMCID: PMC3424585
- DOI: 10.1093/nar/gks610
A high-throughput next-generation sequencing-based method for detecting the mutational fingerprint of carcinogens
Abstract
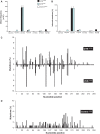
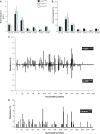
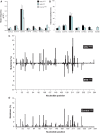
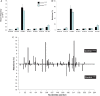
Many carcinogens leave a unique mutational fingerprint in the human genome. These mutational fingerprints manifest as specific types of mutations often clustering at certain genomic loci in tumor genomes from carcinogen-exposed individuals. To develop a high-throughput method for detecting the mutational fingerprint of carcinogens, we have devised a cost-, time- and labor-effective strategy, in which the widely used transgenic Big Blue mouse mutation detection assay is made compatible with the Roche/454 Genome Sequencer FLX Titanium next-generation sequencing technology. As proof of principle, we have used this novel method to establish the mutational fingerprints of three prominent carcinogens with varying mutagenic potencies, including sunlight ultraviolet radiation, 4-aminobiphenyl and secondhand smoke that are known to be strong, moderate and weak mutagens, respectively. For verification purposes, we have compared the mutational fingerprints of these carcinogens obtained by our newly developed method with those obtained by parallel analyses using the conventional low-throughput approach, that is, standard mutation detection assay followed by direct DNA sequencing using a capillary DNA sequencer. We demonstrate that this high-throughput next-generation sequencing-based method is highly specific and sensitive to detect the mutational fingerprints of the tested carcinogens. The method is reproducible, and its accuracy is comparable with that of the currently available low-throughput method. In conclusion, this novel method has the potential to move the field of carcinogenesis forward by allowing high-throughput analysis of mutations induced by endogenous and/or exogenous genotoxic agents.
Figures
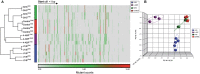
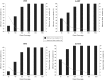
References
-
- DeMarini DM. Mutation spectra of complex mixtures. Mutat. Res. 1998;411:11–18. - PubMed
-
- Hussain SP, Harris CC. Molecular epidemiology of human cancer: contribution of mutation spectra studies of tumor suppressor genes. Cancer Res. 1998;58:4023–4037. - PubMed
-
- Besaratinia A, Pfeifer GP. Investigating human cancer etiology by DNA lesion footprinting and mutagenicity analysis. Carcinogenesis. 2006;27:1526–1537. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

