DNA methylome signature in rheumatoid arthritis
- PMID: 22736089
- PMCID: PMC3549371
- DOI: 10.1136/annrheumdis-2012-201526
DNA methylome signature in rheumatoid arthritis
Abstract
Objectives: Epigenetics can influence disease susceptibility and severity. While DNA methylation of individual genes has been explored in autoimmunity, no unbiased systematic analyses have been reported. Therefore, a genome-wide evaluation of DNA methylation loci in fibroblast-like synoviocytes (FLS) isolated from the site of disease in rheumatoid arthritis (RA) was performed.
Methods: Genomic DNA was isolated from six RA and five osteoarthritis (OA) FLS lines and evaluated using the Illumina HumanMethylation450 chip. Cluster analysis of data was performed and corrected using Benjamini-Hochberg adjustment for multiple comparisons. Methylation was confirmed by pyrosequencing and gene expression was determined by qPCR. Pathway analysis was performed using the Kyoto Encyclopedia of Genes and Genomes.
Results: RA and control FLS segregated based on DNA methylation, with 1859 differentially methylated loci. Hypomethylated loci were identified in key genes relevant to RA, such as CHI3L1, CASP1, STAT3, MAP3K5, MEFV and WISP3. Hypermethylation was also observed, including TGFBR2 and FOXO1. Hypomethylation of individual genes was associated with increased gene expression. Grouped analysis identified 207 hypermethylated or hypomethylated genes with multiple differentially methylated loci, including COL1A1, MEFV and TNF. Hypomethylation was increased in multiple pathways related to cell migration, including focal adhesion, cell adhesion, transendothelial migration and extracellular matrix interactions. Confirmatory studies with OA and normal FLS also demonstrated segregation of RA from control FLS based on methylation pattern.
Conclusions: Differentially methylated genes could alter FLS gene expression and contribute to the pathogenesis of RA. DNA methylation of critical genes suggests that RA FLS are imprinted and implicate epigenetic contributions to inflammatory arthritis.
Figures
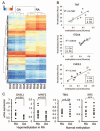

References
-
- Silman AJ, MacGregor AJ, Thomson W, et al. Twin concordance rates for rheumatoid arthritis: results from a nationwide study. Br J Rheumatol. 1993;32:903–7. - PubMed
-
- Jarvinen P, Aho K. Twin studies in rheumatic diseases. Semin Arthritis Rheum. 1994;24:19–28. - PubMed
-
- Issa JP, Ottaviano YL, Celano P, et al. Methylation of the oestrogen receptor CpG island links ageing and neoplasia in human colon. Nat Genet. 1994;7:536–40. - PubMed
-
- Esteller M. CpG island hypermethylation and tumor suppressor genes: a booming present, a brighter future. Oncogene. 2002;21:5427–40. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

