Mammalian alteration/deficiency in activation 3 (Ada3) is essential for embryonic development and cell cycle progression
- PMID: 22736770
- PMCID: PMC3436190
- DOI: 10.1074/jbc.M112.378901
Mammalian alteration/deficiency in activation 3 (Ada3) is essential for embryonic development and cell cycle progression
Abstract
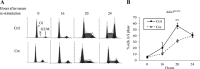
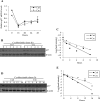
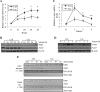
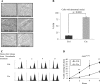
Ada3 protein is an essential component of histone acetyl transferase containing coactivator complexes conserved from yeast to human. We show here that germline deletion of Ada3 in mouse is embryonic lethal, and adenovirus-Cre mediated conditional deletion of Ada3 in Ada3(FL/FL) mouse embryonic fibroblasts leads to a severe proliferation defect which was rescued by ectopic expression of human Ada3. A delay in G(1) to S phase of cell cycle was also seen that was due to accumulation of Cdk inhibitor p27 which was an indirect effect of c-myc gene transcription control by Ada3. We further showed that this defect could be partially reverted by knocking down p27. Additionally, drastic changes in global histone acetylation and changes in global gene expression were observed in microarray analyses upon loss of Ada3. Lastly, formation of abnormal nuclei, mitotic defects and delay in G(2)/M to G(1) transition was seen in Ada3 deleted cells. Taken together, we provide evidence for a critical role of Ada3 in embryogenesis and cell cycle progression as an essential component of HAT complex.
Figures





References
-
- Schafer K. A. (1998) The cell cycle: a review. Vet. Pathol 35, 461–478 - PubMed
-
- Li B., Carey M., Workman J. L. (2007) The role of chromatin during transcription. Cell 128, 707–719 - PubMed
-
- Luger K., Mäder A. W., Richmond R. K., Sargent D. F., Richmond T. J. (1997) Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389, 251–260 - PubMed
-
- Kouzarides T. (2007) Chromatin modifications and their function. Cell 128, 693–705 - PubMed
-
- Strahl B. D., Allis C. D. (2000) The language of covalent histone modifications. Nature 403, 41–45 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- R01 CA087986/CA/NCI NIH HHS/United States
- CA105489/CA/NCI NIH HHS/United States
- U01 CA151806/CA/NCI NIH HHS/United States
- CA116552/CA/NCI NIH HHS/United States
- R01 CA116552/CA/NCI NIH HHS/United States
- P20 RR016469/RR/NCRR NIH HHS/United States
- CA99163/CA/NCI NIH HHS/United States
- CA144027/CA/NCI NIH HHS/United States
- 5U01CA151806-02/CA/NCI NIH HHS/United States
- CA87986/CA/NCI NIH HHS/United States
- R01 CA096844/CA/NCI NIH HHS/United States
- P20 GM103427/GM/NIGMS NIH HHS/United States
- CA96844/CA/NCI NIH HHS/United States
- 8P20GM103427/GM/NIGMS NIH HHS/United States
- R01 CA144027/CA/NCI NIH HHS/United States
- R01 CA105489/CA/NCI NIH HHS/United States
- R01 CA099163/CA/NCI NIH HHS/United States
- 5P20RR016469/RR/NCRR NIH HHS/United States
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

