Functional analysis of the Autographa californica multiple nucleopolyhedrovirus GP64 terminal fusion loops and interactions with membranes
- PMID: 22740400
- PMCID: PMC3446601
- DOI: 10.1128/JVI.00813-12
Functional analysis of the Autographa californica multiple nucleopolyhedrovirus GP64 terminal fusion loops and interactions with membranes
Abstract
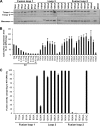
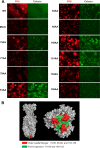
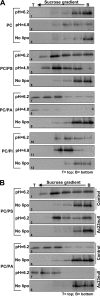
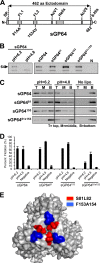
The Autographa californica multiple nucleopolyhedrovirus (AcMNPV) glycoprotein GP64 is the major envelope protein of the budded virus (BV). GP64 is a class III fusion protein that mediates BV attachment to the cell surface and low-pH-triggered membrane fusion between the BV envelope and the endosome membrane during entry. Class III fusion proteins contain terminal looped structures that are believed to interact with membranes. To examine the functions of 3 loops found at the apex of the GP64 postfusion structure, we generated 2-alanine substitutions that scanned the two so-called fusion loops (loop 1 and loop 2) plus an adjacent loop structure (loop 3) that is closely attached to loop 2 and is also found at the apex of the GP64 postfusion structure. We identified essential residues from Y75 to T86 (loop 1) and N149 to H156 (loop 2) that are required for fusion activity, but no essential residues in loop 3. Further analysis revealed that critical fusion loop residues fall within two groups that are associated with either membrane merger (hemifusion) or fusion pore expansion. We next examined the interactions of soluble GP64 proteins and BV with membranes composed of various phospholipids. BV interacted directly with small unilamellar vesicles (SUVs) comprised of phospholipids phosphatidylcholine and phosphatidic acid (PC/PA) or phosphatidylcholine and phosphatidylserine (PC/PS) under neutral and acidic pH. We also examined the interactions of soluble GP64 constructs containing substitutions of the most hydrophobic residues within each of the two fusion loops. We found that a 2-residue substitution in either single loop (loop 1 [positions 81 and 82] or loop 2 [positions 153 and 154]) was not sufficient to substantially reduce the GP64-liposome interaction, but the same substitutions in both fusion loops severely reduced the GP64-liposome association at neutral pH. These results suggest that critical hydrophobic residues in both fusion loops may be involved in the interaction of GP64 with host cellular membranes and direct GP64-membrane interactions may represent a receptor-binding step prior to a low-pH-triggered conformational change.
Figures

Similar articles
-
Critical Residues and Contacts within Domain IV of Autographa californica Multiple Nucleopolyhedrovirus GP64 Contribute to Its Refolding during Membrane Fusion.J Virol. 2020 Sep 15;94(19):e01105-20. doi: 10.1128/JVI.01105-20. Print 2020 Sep 15. J Virol. 2020. PMID: 32699096 Free PMC article.
-
The pre-transmembrane domain of the Autographa californica multicapsid nucleopolyhedrovirus GP64 protein is critical for membrane fusion and virus infectivity.J Virol. 2009 Nov;83(21):10993-1004. doi: 10.1128/JVI.01085-09. Epub 2009 Aug 19. J Virol. 2009. PMID: 19692475 Free PMC article.
-
Autographa californica multiple nucleopolyhedrovirus GP64 protein: roles of histidine residues in triggering membrane fusion and fusion pore expansion.J Virol. 2011 Dec;85(23):12492-504. doi: 10.1128/JVI.05153-11. Epub 2011 Sep 21. J Virol. 2011. PMID: 21937651 Free PMC article.
-
me53 encoded by Autographa californica multiple nucleopolyhedrovirus: from mechanism to function.Virus Genes. 2023 Apr;59(2):188-194. doi: 10.1007/s11262-022-01943-3. Epub 2022 Oct 13. Virus Genes. 2023. PMID: 36229721 Review.
-
Class II fusion proteins.Adv Exp Med Biol. 2013;790:150-66. doi: 10.1007/978-1-4614-7651-1_8. Adv Exp Med Biol. 2013. PMID: 23884590 Free PMC article. Review.
Cited by
-
Structural transition of GP64 triggered by a pH-sensitive multi-histidine switch.Nat Commun. 2024 Sep 3;15(1):7668. doi: 10.1038/s41467-024-51799-4. Nat Commun. 2024. PMID: 39227374 Free PMC article.
-
BmREEPa Is a Novel Gene that Facilitates BmNPV Entry into Silkworm Cells.PLoS One. 2015 Dec 14;10(12):e0144575. doi: 10.1371/journal.pone.0144575. eCollection 2015. PLoS One. 2015. PMID: 26656276 Free PMC article.
-
Critical Residues and Contacts within Domain IV of Autographa californica Multiple Nucleopolyhedrovirus GP64 Contribute to Its Refolding during Membrane Fusion.J Virol. 2020 Sep 15;94(19):e01105-20. doi: 10.1128/JVI.01105-20. Print 2020 Sep 15. J Virol. 2020. PMID: 32699096 Free PMC article.
-
Microscopic Observation of Membrane Fusion between Giant Liposomes and Baculovirus Budded Viruses Activated by the Release of a Caged Proton.Membranes (Basel). 2023 May 11;13(5):507. doi: 10.3390/membranes13050507. Membranes (Basel). 2023. PMID: 37233568 Free PMC article.
-
Novel GP64 envelope variants for improved delivery to human airway epithelial cells.Gene Ther. 2017 Oct;24(10):674-679. doi: 10.1038/gt.2017.78. Epub 2017 Sep 7. Gene Ther. 2017. PMID: 28880020 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

