Structural basis for the reaction mechanism of S-carbamoylation of HypE by HypF in the maturation of [NiFe]-hydrogenases
- PMID: 22740694
- PMCID: PMC3436534
- DOI: 10.1074/jbc.M112.387134
Structural basis for the reaction mechanism of S-carbamoylation of HypE by HypF in the maturation of [NiFe]-hydrogenases
Abstract
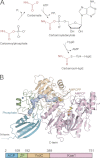
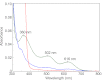
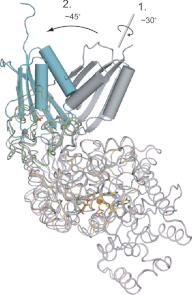
As a remarkable structural feature of hydrogenase active sites, [NiFe]-hydrogenases harbor one carbonyl and two cyano ligands, where HypE and HypF are involved in the biosynthesis of the nitrile group as a precursor of the cyano groups. HypF catalyzes S-carbamoylation of the C-terminal cysteine of HypE via three steps using carbamoylphosphate and ATP, producing two unstable intermediates: carbamate and carbamoyladenylate. Although the crystal structures of intact HypE homodimers and partial HypF have been reported, it remains unclear how the consecutive reactions occur without the loss of unstable intermediates during the proposed reaction scheme. Here we report the crystal structures of full-length HypF both alone and in complex with HypE at resolutions of 2.0 and 2.6 Å, respectively. Three catalytic sites of the structures of the HypF nucleotide- and phosphate-bound forms have been identified, with each site connected via channels inside the protein. This finding suggests that the first two consecutive reactions occur without the release of carbamate or carbamoyladenylate from the enzyme. The structure of HypF in complex with HypE revealed that HypF can associate with HypE without disturbing its homodimeric interaction and that the binding manner allows the C-terminal Cys-351 of HypE to access the S-carbamoylation active site in HypF, suggesting that the third step can also proceed without the release of carbamoyladenylate. A comparison of the structure of HypF with the recently reported structures of O-carbamoyltransferase revealed different reaction mechanisms for carbamoyladenylate synthesis and a similar reaction mechanism for carbamoyltransfer to produce the carbamoyl-HypE molecule.
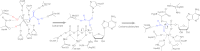
Figures

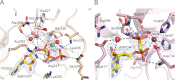

Similar articles
-
Analysis of the transcarbamoylation-dehydration reaction catalyzed by the hydrogenase maturation proteins HypF and HypE.Eur J Biochem. 2004 Aug;271(16):3428-36. doi: 10.1111/j.1432-1033.2004.04280.x. Eur J Biochem. 2004. PMID: 15291820
-
Structural Insight into [NiFe] Hydrogenase Maturation by Transient Complexes between Hyp Proteins.Acc Chem Res. 2020 Apr 21;53(4):875-886. doi: 10.1021/acs.accounts.0c00022. Epub 2020 Mar 31. Acc Chem Res. 2020. PMID: 32227866 Review.
-
Taming of a poison: biosynthesis of the NiFe-hydrogenase cyanide ligands.Science. 2003 Feb 14;299(5609):1067-70. doi: 10.1126/science.1080972. Science. 2003. PMID: 12586941
-
Structure of [NiFe] hydrogenase maturation protein HypE from Escherichia coli and its interaction with HypF.J Bacteriol. 2008 Feb;190(4):1447-58. doi: 10.1128/JB.01610-07. Epub 2007 Dec 7. J Bacteriol. 2008. PMID: 18065529 Free PMC article.
-
Maturation of [NiFe]-hydrogenases in Escherichia coli.Biometals. 2007 Jun;20(3-4):565-78. doi: 10.1007/s10534-006-9048-5. Epub 2007 Jan 11. Biometals. 2007. PMID: 17216401 Review.
Cited by
-
Endowing homodimeric carbamoyltransferase GdmN with iterative functions through structural characterization and mechanistic studies.Nat Commun. 2022 Nov 3;13(1):6617. doi: 10.1038/s41467-022-34387-2. Nat Commun. 2022. PMID: 36329057 Free PMC article.
-
Structure-function analysis of an ancient TsaD-TsaC-SUA5-TcdA modular enzyme reveals a prototype of tRNA t6A and ct6A synthetases.Nucleic Acids Res. 2023 Sep 8;51(16):8711-8729. doi: 10.1093/nar/gkad587. Nucleic Acids Res. 2023. PMID: 37427786 Free PMC article.
-
Biosynthesis and assembly of hydrogenase [NiFe]-cofactor: recent advances and perspectives.Metallomics. 2025 Jun 3;17(6):mfaf015. doi: 10.1093/mtomcs/mfaf015. Metallomics. 2025. PMID: 40411749 Free PMC article. Review.
-
Structural characterization of HypX responsible for CO biosynthesis in the maturation of NiFe-hydrogenase.Commun Biol. 2019 Oct 18;2:385. doi: 10.1038/s42003-019-0631-z. eCollection 2019. Commun Biol. 2019. PMID: 31646188 Free PMC article.
-
Efficient anaerobic consumption of D-xylose by E. coli BL21(DE3) via xylR adaptive mutation.BMC Microbiol. 2021 Dec 6;21(1):332. doi: 10.1186/s12866-021-02395-9. BMC Microbiol. 2021. PMID: 34872501 Free PMC article.
References
-
- Vignais P. M., Billoud B. (2007) Occurrence, classification, and biological function of hydrogenases: an overview. Chem. Rev. 107, 4206–4272 - PubMed
-
- Fontecilla-Camps J. C., Volbeda A., Cavazza C., Nicolet Y. (2007) Structure/function relationships of [NiFe]- and [FeFe]-hydrogenases. Chem. Rev. 107, 4273–4303 - PubMed
-
- Shima S., Thauer R. K. (2007) A third type of hydrogenase catalyzing H2 activation. Chem. Rec. 7, 37–46 - PubMed
-
- Reissmann S., Hochleitner E., Wang H., Paschos A., Lottspeich F., Glass R. S., Böck A. (2003) Taming of a poison: biosynthesis of the NiFe-hydrogenase cyanide ligands. Science 299, 1067–1070 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Research Materials

