Catalytic zinc site and mechanism of the metalloenzyme PR-AMP cyclohydrolase
- PMID: 22741521
- PMCID: PMC3427388
- DOI: 10.1021/bi300391m
Catalytic zinc site and mechanism of the metalloenzyme PR-AMP cyclohydrolase
Abstract
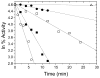
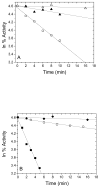
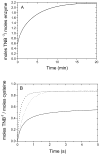
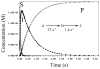
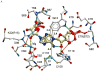
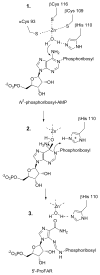
The enzyme N(1)-(5'-phosphoribosyl) adenosine-5'-monophosphate cyclohydrolase (PR-AMP cyclohydrolase) is a Zn(2+) metalloprotein encoded by the hisI gene. It catalyzes the third step of histidine biosynthesis, an uncommon ring-opening of a purine heterocycle for use in primary metabolism. A three-dimensional structure of the enzyme from Methanobacterium thermoautotrophicum has revealed that three conserved cysteine residues occur at the dimer interface and likely form the catalytic site. To investigate the functions of these cysteines in the enzyme from Methanococcus vannielii, a series of biochemical studies were pursued to test the basic hypothesis regarding their roles in catalysis. Inactivation of the enzyme activity by methyl methane thiosulfonate (MMTS) or 5,5'-dithiobis(2-nitrobenzoic acid) (DTNB) also compromised the Zn(2+) binding properties of the protein inducing loss of up to 90% of the metal. Overall reaction stoichiometry and the potassium cyanide (KCN) induced cleavage of the protein suggested that all three cysteines were modified in the process. The enzyme was protected from DTNB-induced inactivation by inclusion of the substrate N(1)-(5'-phosphoribosyl)adenosine 5'-monophosphate; (PR-AMP), while Mg(2+), a metal required for catalytic activity, enhanced the rate of inactivation. Site-directed mutations of the conserved C93, C109, C116 and the double mutant C109/C116 were prepared and analyzed for catalytic activity, Zn(2+) content, and reactivity with DTNB. Substitution of alanine for each of the conserved cysteines showed no measurable catalytic activity, and only the C116A was still capable of binding Zn(2+). Reactions of DTNB with the C109A/C116A double mutant showed that C93 is completely modified within 0.5 s. A model consistent with these data involves a DTNB-induced mixed disulfide linkage between C93 and C109 or C116, followed by ejection of the active site Zn(2+) and provides further evidence that the Zn(2+) coordination site involves the three conserved cysteine residues. The C93 reactivity is modulated by the presence of the Zn(2+) and Mg(2+) and substantiates the role of this residue as a metal ligand. In addition, Mg(2+) ligand binding site(s) indicated by the structural analysis were probed by site-directed mutagenesis of three key aspartate residues flanking the conserved C93 which were shown to have a functional impact on catalysis, cysteine activation, and metal (zinc) binding capacity. The unique amino acid sequence, the dynamic properties of the cysteine ligands involved in Zn(2+) coordination, and the requirement for a second metal (Mg(2+)) are discussed in the context of their roles in catalysis. The results are consistent with a Zn(2+)-mediated activation of H(2)O mechanism involving histidine as a general base that has features similar to but distinct from those of previously characterized purine and pyrimidine deaminases.
Figures



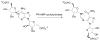

References
-
- Lee D-S, Burd H, Liu J, Almaas E, Wiest O, Barabási A-L, Oltvai ZN, Kapatral V. Comparative Genome-Scale Metabolic Reconstruction and Flux Balance Analysis of Multiple Staphylococcus aureus Genomes Identify Novel Antimicrobial Drug Targets. Journal of Bacteriology. 2009;191:4015–4024. - PMC - PubMed
-
- D'Ordine RL, Klem TJ, Davisson VJ. N1-(5'-phosphoribosyl)adenosine -5'-Monophosphate Cyclohydrolase: Purification and Characterization of A Unique Metalloenzyme. Biochemistry. 1999;38:1537–1546. - PubMed
-
- Jones W, Kurz LC, Wolfenden R. Transition-State Stabilization by Adenosine Deaminase:16-Addition of Water to Purine Ribonucleoside The Enzyme's Affinity for 6-Hydroxy-16-dihydropurine Ribonucleoside and the Effective Concentration of Substrate Water at the Active Site. Biochemistry. 1989;28:1242–1247. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

