+TIPs: SxIPping along microtubule ends
- PMID: 22748381
- PMCID: PMC3408562
- DOI: 10.1016/j.tcb.2012.05.005
+TIPs: SxIPping along microtubule ends
Abstract
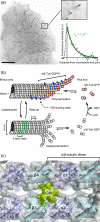
+TIPs are a heterogeneous class of proteins that specifically bind to growing microtubule ends. Because dynamic microtubules are essential for many intracellular processes, +TIPs play important roles in regulating microtubule dynamics and microtubule interactions with other intracellular structures. End-binding proteins (EBs) recognize a structural cap at growing microtubule ends, and have emerged as central adaptors that mediate microtubule plus-end tracking of potentially all other +TIPs. The majority of these +TIPs bind to EBs through a short hydrophobic (S/T)x(I/L)P sequence motif (SxIP) and surrounding electrostatic interactions. These recent discoveries have resulted in a rapid expansion of the number of possible +TIPs. In this review, we outline our current understanding of the molecular mechanism of plus-end tracking and provide an overview of SxIP-recruited +TIPs.
Copyright © 2012 Elsevier Ltd. All rights reserved.
Figures

References
-
- Perez F, et al. CLIP-170 highlights growing microtubule ends in vivo. Cell. 1999;96:517–527. - PubMed
-
- Mimori-Kiyosue Y, et al. The dynamic behavior of the APC-binding protein EB1 on the distal ends of microtubules. Curr. Biol. 2000;10:865–868. - PubMed
-
- Galjart N. Plus-end-tracking proteins and their interactions at microtubule ends. Curr. Biol. 2010;20:R528–R537. - PubMed
-
- Akhmanova A, Steinmetz MO. Tracking the ends: a dynamic protein network controls the fate of microtubule tips. Nat. Rev. Mol. Cell Biol. 2008;9:309–322. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

