Fatty acid biosynthesis in Pseudomonas aeruginosa is initiated by the FabY class of β-ketoacyl acyl carrier protein synthases
- PMID: 22753059
- PMCID: PMC3457228
- DOI: 10.1128/JB.00792-12
Fatty acid biosynthesis in Pseudomonas aeruginosa is initiated by the FabY class of β-ketoacyl acyl carrier protein synthases
Abstract
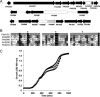
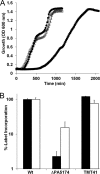
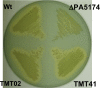
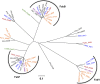
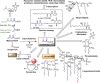
The prototypical type II fatty acid synthesis (FAS) pathway in bacteria utilizes two distinct classes of β-ketoacyl synthase (KAS) domains to assemble long-chain fatty acids, the KASIII domain for initiation and the KASI/II domain for elongation. The central role of FAS in bacterial viability and virulence has stimulated significant effort toward developing KAS inhibitors, particularly against the KASIII domain of the β-acetoacetyl-acyl carrier protein (ACP) synthase FabH. Herein, we show that the opportunistic pathogen Pseudomonas aeruginosa does not utilize a FabH ortholog but rather a new class of divergent KAS I/II enzymes to initiate the FAS pathway. When a P. aeruginosa cosmid library was used to rescue growth in a fabH downregulated strain of Escherichia coli, a single unannotated open reading frame, PA5174, complemented fabH depletion. While deletion of all four KASIII domain-encoding genes in the same P. aeruginosa strain resulted in a wild-type growth phenotype, deletion of PA5174 alone specifically attenuated growth due to a defect in de novo FAS. Siderophore secretion and quorum-sensing signaling, particularly in the rhl and Pseudomonas quinolone signal (PQS) systems, was significantly muted in the absence of PA5174. The defect could be repaired by intergeneric complementation with E. coli fabH. Characterization of recombinant PA5174 confirmed a preference for short-chain acyl coenzyme A (acyl-CoA) substrates, supporting the identification of PA5174 as the predominant enzyme catalyzing the condensation of acetyl coenzyme A with malonyl-ACP in P. aeruginosa. The identification of the functional role for PA5174 in FAS defines the new FabY class of β-ketoacyl synthase KASI/II domain condensation enzymes.
Figures





Comment in
-
Will the initiator of fatty acid synthesis in Pseudomonas aeruginosa please stand up?J Bacteriol. 2012 Oct;194(19):5159-61. doi: 10.1128/JB.01198-12. Epub 2012 Jul 20. J Bacteriol. 2012. PMID: 22821980 Free PMC article. No abstract available.
References
-
- Alberts AW, Bell RM, Vagelos PR. 1972. Acyl carrier protein. XV. Studies of β-ketoacyl-acyl carrier protein synthetase. J. Biol. Chem. 247:3190–3198 - PubMed
-
- Alberts AW, Majerus PW, Talamo B, Vagelos PR. 1964. Acyl-carrier protein. II. Intermediary reactions of fatty acid synthesis. Biochemistry 3:1563–1571 - PubMed
-
- Bergler H, et al. 1994. Protein EnvM is the NADH-dependent enoyl-ACP reductase (FabI) of Escherichia coli. J. Biol. Chem. 269:5493–5496 - PubMed
-
- Bredenbruch F, Geffers R, Nimtz M, Buer J, Haussler S. 2006. The Pseudomonas aeruginosa quinolone signal (PQS) has an iron-chelating activity. Environ. Microbiol. 8:1318–1329 - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

