Heat shock protein 90α (HSP90α), a substrate and chaperone of DNA-PK necessary for the apoptotic response
- PMID: 22753480
- PMCID: PMC3420188
- DOI: 10.1073/pnas.1203617109
Heat shock protein 90α (HSP90α), a substrate and chaperone of DNA-PK necessary for the apoptotic response
Abstract
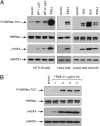
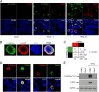
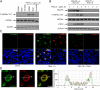
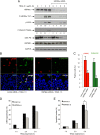
The "apoptotic ring" is characterized by the phosphorylation of histone H2AX at serine 139 (γ-H2AX) by DNA-dependent protein kinase (DNA-PK). The γ-H2AX apoptotic ring differs from the nuclear foci patterns observed in response to DNA-damaging agents. It contains phosphorylated DNA damage response proteins including activated Chk2, activated ATM, and activated DNA-PK itself but lacks MDC1 and 53BP1, which are required to initiate DNA repair. Because DNA-PK can phosphorylate heat shock protein 90α (HSP90α) in biochemical assays, we investigated whether HSP90α is involved in the apoptotic ring. Here we show that HSP90α is phosphorylated by DNA-PK on threonines 5 and 7 early during apoptosis and that both phosphorylated HSP90α and DNA-PK colocalize in the apoptotic ring. We also show that DNA-PK is a client of HSP90α and that HSP90α is required for full DNA-PK activation, γ-H2AX formation, DNA fragmentation, and apoptotic body formation. In contrast, HSP90 inhibition by geldanamycin markedly enhances TRAIL-induced DNA-PK and H2AX activation. Together, our results reveal that HSP90α is a substrate and chaperone of DNA-PK in the apoptotic response. The response of phosphorylated HSP90α to TRAIL and its localization to the γ-H2AX ring represent epigenetic features of apoptosis that offer insights for studying and monitoring nuclear apoptosis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
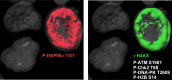
Comment in
-
Safety control for apoptotic irreversibility.Proc Natl Acad Sci U S A. 2012 Aug 7;109(32):12844-5. doi: 10.1073/pnas.1210736109. Epub 2012 Jul 24. Proc Natl Acad Sci U S A. 2012. PMID: 22829668 Free PMC article. No abstract available.
Similar articles
-
DNA-PK phosphorylates histone H2AX during apoptotic DNA fragmentation in mammalian cells.DNA Repair (Amst). 2006 May 10;5(5):575-90. doi: 10.1016/j.dnarep.2006.01.011. Epub 2006 Mar 29. DNA Repair (Amst). 2006. PMID: 16567133
-
Death receptor-induced activation of the Chk2- and histone H2AX-associated DNA damage response pathways.Mol Cell Biol. 2009 Jan;29(1):68-82. doi: 10.1128/MCB.00581-08. Epub 2008 Oct 27. Mol Cell Biol. 2009. PMID: 18955500 Free PMC article.
-
Heat shock protein 90α (Hsp90α) is phosphorylated in response to DNA damage and accumulates in repair foci.J Biol Chem. 2012 Mar 16;287(12):8803-15. doi: 10.1074/jbc.M111.320887. Epub 2012 Jan 23. J Biol Chem. 2012. PMID: 22270370 Free PMC article.
-
Involvement of DNA-PK and ATM in radiation- and heat-induced DNA damage recognition and apoptotic cell death.J Radiat Res. 2010;51(5):493-501. doi: 10.1269/jrr.10039. Epub 2010 Aug 28. J Radiat Res. 2010. PMID: 20814172 Review.
-
The nuclear γ-H2AX apoptotic ring: implications for cancers and autoimmune diseases.Cell Mol Life Sci. 2014 Jun;71(12):2289-97. doi: 10.1007/s00018-013-1555-2. Epub 2014 Jan 22. Cell Mol Life Sci. 2014. PMID: 24448903 Free PMC article. Review.
Cited by
-
ATM is the primary kinase responsible for phosphorylation of Hsp90α after ionizing radiation.Oncotarget. 2016 Dec 13;7(50):82450-82457. doi: 10.18632/oncotarget.12557. Oncotarget. 2016. PMID: 27738310 Free PMC article.
-
Detection and repair of ionizing radiation-induced DNA double strand breaks: new developments in nonhomologous end joining.Int J Radiat Oncol Biol Phys. 2013 Jul 1;86(3):440-9. doi: 10.1016/j.ijrobp.2013.01.011. Epub 2013 Feb 20. Int J Radiat Oncol Biol Phys. 2013. PMID: 23433795 Free PMC article. Review.
-
Silencing of Hsp70 intensifies 6-OHDA-induced apoptosis and Hsp90 upregulation in PC12 cells.J Mol Neurosci. 2015 Jan;55(1):174-183. doi: 10.1007/s12031-014-0298-3. Epub 2014 Apr 13. J Mol Neurosci. 2015. PMID: 24729093
-
Heat shock proteins and DNA repair mechanisms: an updated overview.Cell Stress Chaperones. 2018 May;23(3):303-315. doi: 10.1007/s12192-017-0843-4. Epub 2017 Sep 26. Cell Stress Chaperones. 2018. PMID: 28952019 Free PMC article. Review.
-
Regulation and function of the human HSP90AA1 gene.Gene. 2015 Oct 1;570(1):8-16. doi: 10.1016/j.gene.2015.06.018. Epub 2015 Jun 10. Gene. 2015. PMID: 26071189 Free PMC article. Review.
References
-
- Neckers L. Heat shock protein 90: The cancer chaperone. J Biosci. 2007;32:517–530. - PubMed
-
- Wandinger SK, Richter K, Buchner J. The Hsp90 chaperone machinery. J Biol Chem. 2008;283:18473–18477. - PubMed
-
- Zhao R, et al. Navigating the chaperone network: An integrative map of physical and genetic interactions mediated by the hsp90 chaperone. Cell. 2005;120:715–727. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

