The box C/D sRNP dimeric architecture is conserved across domain Archaea
- PMID: 22753779
- PMCID: PMC3404373
- DOI: 10.1261/rna.033134.112
The box C/D sRNP dimeric architecture is conserved across domain Archaea
Abstract
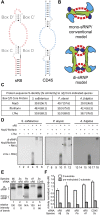
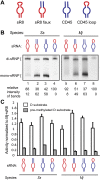
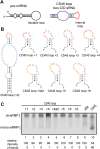
Box C/D small (nucleolar) ribonucleoproteins [s(no)RNPs] catalyze RNA-guided 2'-O-ribose methylation in two of the three domains of life. Recent structural studies have led to a controversy over whether box C/D sRNPs functionally assemble as monomeric or dimeric macromolecules. The archaeal box C/D sRNP from Methanococcus jannaschii (Mj) has been shown by glycerol gradient sedimentation, gel filtration chromatography, native gel analysis, and single-particle electron microscopy (EM) to adopt a di-sRNP architecture, containing four copies of each box C/D core protein and two copies of the Mj sR8 sRNA. Subsequently, investigators used a two-stranded artificial guide sRNA, CD45, to assemble a box C/D sRNP from Sulfolobus solfataricus with a short RNA methylation substrate, yielding a crystal structure of a mono-sRNP. To more closely examine box C/D sRNP architecture, we investigate the role of the omnipresent sRNA loop as a structural determinant of sRNP assembly. We show through sRNA mutagenesis, native gel electrophoresis, and single-particle EM that a di-sRNP is the near exclusive architecture obtained when reconstituting box C/D sRNPs with natural or artificial sRNAs containing an internal loop. Our results span three distantly related archaeal species--Sulfolobus solfataricus, Pyrococcus abyssi, and Archaeoglobus fulgidus--indicating that the di-sRNP architecture is broadly conserved across the entire archaeal domain.
Figures




Similar articles
-
Assembly of the archaeal box C/D sRNP can occur via alternative pathways and requires temperature-facilitated sRNA remodeling.J Mol Biol. 2006 Oct 6;362(5):1025-42. doi: 10.1016/j.jmb.2006.07.091. Epub 2006 Aug 4. J Mol Biol. 2006. PMID: 16949610
-
The coiled-coil domain of the Nop56/58 core protein is dispensable for sRNP assembly but is critical for archaeal box C/D sRNP-guided nucleotide methylation.RNA. 2006 Jun;12(6):1092-103. doi: 10.1261/rna.2230106. Epub 2006 Apr 6. RNA. 2006. PMID: 16601205 Free PMC article.
-
Dissecting the role of conserved box C/D sRNA sequences in di-sRNP assembly and function.Nucleic Acids Res. 2010 Dec;38(22):8295-305. doi: 10.1093/nar/gkq690. Epub 2010 Aug 6. Nucleic Acids Res. 2010. PMID: 20693534 Free PMC article.
-
The structure and function of small nucleolar ribonucleoproteins.Nucleic Acids Res. 2007;35(5):1452-64. doi: 10.1093/nar/gkl1172. Epub 2007 Feb 6. Nucleic Acids Res. 2007. PMID: 17284456 Free PMC article. Review.
-
Small non-coding RNAs in Archaea.Curr Opin Microbiol. 2005 Dec;8(6):685-94. doi: 10.1016/j.mib.2005.10.013. Epub 2005 Oct 26. Curr Opin Microbiol. 2005. PMID: 16256421 Review.
Cited by
-
Ribonucleoproteins in archaeal pre-rRNA processing and modification.Archaea. 2013;2013:614735. doi: 10.1155/2013/614735. Epub 2013 Mar 10. Archaea. 2013. PMID: 23554567 Free PMC article. Review.
-
Production of nascent ribosome precursors within the nucleolar microenvironment of Saccharomyces cerevisiae.Genetics. 2022 Jul 4;221(3):iyac070. doi: 10.1093/genetics/iyac070. Genetics. 2022. PMID: 35657327 Free PMC article.
-
Ribosome biogenesis in the yeast Saccharomyces cerevisiae.Genetics. 2013 Nov;195(3):643-81. doi: 10.1534/genetics.113.153197. Genetics. 2013. PMID: 24190922 Free PMC article. Review.
-
The structure of the box C/D enzyme reveals regulation of RNA methylation.Nature. 2013 Oct 24;502(7472):519-23. doi: 10.1038/nature12581. Epub 2013 Oct 13. Nature. 2013. PMID: 24121435
-
Co-expression and co-purification of archaeal and eukaryal box C/D RNPs.PLoS One. 2014 Jul 31;9(7):e103096. doi: 10.1371/journal.pone.0103096. eCollection 2014. PLoS One. 2014. PMID: 25078083 Free PMC article.
References
-
- Abramovitz DL, Pyle AM 1997. Remarkable morphological variability of a common RNA folding motif: The GNRA tetraloop–receptor interaction. J Mol Biol 266: 493–506 - PubMed
-
- Aittaleb M, Rashid R, Chen Q, Palmer JR, Daniels CJ, Li H 2003. Structure and function of archaeal box C/D sRNP core proteins. Nat Struct Biol 10: 256–263 - PubMed
-
- Aittaleb M, Visone T, Fenley MO, Li H 2004. Structural and thermodynamic evidence for a stabilizing role of Nop5p in S-adenosyl-L-methionine binding to fibrillarin. J Biol Chem 279: 41822–41829 - PubMed
-
- Balakin AG, Smith L, Fournier MJ 1996. The RNA world of the nucleolus: Two major families of small RNAs defined by different box elements with related functions. Cell 86: 823–834 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
