Mathematical and statistical modeling in cancer systems biology
- PMID: 22754537
- PMCID: PMC3385354
- DOI: 10.3389/fphys.2012.00227
Mathematical and statistical modeling in cancer systems biology
Abstract
Cancer is a major health problem with high mortality rates. In the post-genome era, investigators have access to massive amounts of rapidly accumulating high-throughput data in publicly available databases, some of which are exclusively devoted to housing Cancer data. However, data interpretation efforts have not kept pace with data collection, and gained knowledge is not necessarily translating into better diagnoses and treatments. A fundamental problem is to integrate and interpret data to further our understanding in Cancer Systems Biology. Viewing cancer as a network provides insights into the complex mechanisms underlying the disease. Mathematical and statistical models provide an avenue for cancer network modeling. In this article, we review two widely used modeling paradigms: deterministic metabolic models and statistical graphical models. The strength of these approaches lies in their flexibility and predictive power. Once a model has been validated, it can be used to make predictions and generate hypotheses. We describe a number of diverse applications to Cancer Biology, including, the system-wide effects of drug-treatments, disease prognosis, tumor classification, forecasting treatment outcomes, and survival predictions.
Keywords: ODEs; cancer; dynamic; graphical models; high-throughput data; metabolism; steady-state.
Figures
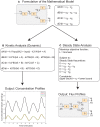

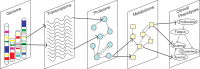
References
-
- Astsaturov I., Ratushny V., Sukhanova A., Einarson M., Bagnyukova T., Zhou Y., Devarajan K., Silverman J., Tikhmyanova N., Skobeleva N., Pecherskaya A., Nasto R., Sharma C., Jablonski S., Serebriiskii I., Weiner L., Golemis E. (2010). Synthetic lethal screen of an egfr-centered network to improve targeted therapies. Sci. Signal. 3, ra67. 10.1126/scisignal.2001083 - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources

