The porcine antibody repertoire: variations on the textbook theme
- PMID: 22754553
- PMCID: PMC3384076
- DOI: 10.3389/fimmu.2012.00153
The porcine antibody repertoire: variations on the textbook theme
Abstract
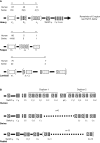
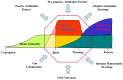
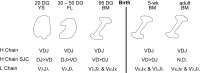
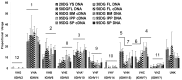
The genes encoding the heavy and light chains of swine antibodies are organized in the same manner as in other eutherian mammals. There are ∼30 VH genes, two functional DH genes and one functional JH gene, 14-60 Vκ genes, 5 Jκ segments, 12-13 functional Vλ genes, and two functional Jλ genes. The heavy chain constant regions encode the same repertoire of isotypes common to other eutherian mammals. The piglet models offers advantage over rodent models since the fetal repertoire develops without maternal influences and the precocial nature of their multiple offspring allows the experimenter to control the influences of environmental and maternal factors on repertoire development postnatally. B cell lymphogenesis in swine begins in the fetal yolk sac at 20 days of gestation (DG), moves to the fetal liver at 30 DG and eventually to the bone marrow which dominates until birth (114 DG) and to at least 5 weeks postpartum. There is no evidence that the ileal Peyers patches are a site of B cell lymphogenesis or are required for B cell maintenance. Unlike rodents and humans, light chain rearrangement begins first in the lambda locus; kappa rearrangements are not seen until late gestation. Dissimilar to lab rodents and more in the direction of the rabbit, swine utilize a small number of VH genes to form >90% of their pre-immune repertoire. Diversification in response to environmental antigen does not alter this pattern and is achieved by somatic hypermutation (SHM) of the same small number of VH genes. The situation for light chains is less well studied, but certain Vκ and Jκ and Vλ and Jλ are dominant in transcripts and in contrast to rearranged heavy chains, there is little junctional diversity, less SHM, and mutations are not concentrated in CDR regions. The transcribed and secreted pre-immune antibodies of the fetus include mainly IgM, IgA, and IgG3; this last isotype may provide a type of first responder mucosal immunity. Development of functional adaptive immunity is dependent on bacterial MAMPs or MAMPs provided by viral infections, indicating the importance of innate immunity for development of adaptive immunity. The structural analysis of Ig genes of this species indicate that especially the VH and Cγ gene are the result of tandem gene duplication in the context of genomic gene conversion. Since only a few of these duplicated VH genes substantially contribute to the antibody repertoire, polygeny may be a vestige from a time before somatic processes became prominently evolved to generate the antibody repertoire. In swine we believe such duplications within the genome have very limited functional significance and their occurrence is therefore overrated.
Keywords: antibody repertoire; development; gene duplication; swine.
Figures


References
-
- Acha-Orbea H., Finke D., Attinger A., Schmid S., Wehrli N., Vacheron S., Xenarios I., Scarpellino L., Toellner K. M., MacLennan I. C., Lutter S. A. (1999). Interplay between mouse mammary tumor virus and the cellular and humoral immune response. Immunol. Rev. 168, 287–303 10.1111/j.1600-065X.1999.tb01299.x - DOI - PubMed
-
- Brambell F. W. R. (1970). The Transmission of Passive immunity from Mother to Young. Amsterdam: North-Holland Publishing Company
LinkOut - more resources
Full Text Sources
Miscellaneous

