Genome plasticity and systems evolution in Streptomyces
- PMID: 22759432
- PMCID: PMC3388780
- DOI: 10.1186/1471-2105-13-S10-S8
Genome plasticity and systems evolution in Streptomyces
Abstract
Background: Streptomycetes are filamentous soil-dwelling bacteria. They are best known as the producers of a great variety of natural products such as antibiotics, antifungals, antiparasitics, and anticancer agents and the decomposers of organic substances for carbon recycling. They are also model organisms for the studies of gene regulatory networks, morphological differentiation, and stress response. The availability of sets of genomes from closely related Streptomyces strains makes it possible to assess the mechanisms underlying genome plasticity and systems adaptation.
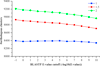
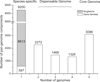
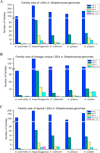
Results: We present the results of a comprehensive analysis of the genomes of five Streptomyces species with distinct phenotypes. These streptomycetes have a pan-genome comprised of 17,362 orthologous families which includes 3,096 components in the core genome, 5,066 components in the dispensable genome, and 9,200 components that are uniquely present in only one species. The core genome makes up about 33%-45% of each genome repertoire. It contains important genes for Streptomyces biology including those involved in gene regulation, secretion, secondary metabolism and morphological differentiation. Abundant duplicate genes have been identified, with 4%-11% of the whole genomes composed of lineage-specific expansions (LSEs), suggesting that frequent gene duplication or lateral gene transfer events play a role in shaping the genome diversification within this genus. Two patterns of expansion, single gene expansion and chromosome block expansion are observed, representing different scales of duplication.
Conclusions: Our results provide a catalog of genome components and their potential functional roles in gene regulatory networks and metabolic networks. The core genome components reveal the minimum requirement for streptomycetes to sustain a successful lifecycle in the soil environment, reflecting the effects of both genome evolution and environmental stress acting upon the expressed phenotypes. A better understanding of the LSE gene families will, on the other hand, bring a wealth of new insights into the mechanisms underlying strain-specific phenotypes, such as the production of novel antibiotics, pathogenesis, and adaptive response to environmental challenges.
Figures


Similar articles
-
Genomic and systems evolution in Vibrionaceae species.BMC Genomics. 2009 Jul 7;10 Suppl 1(Suppl 1):S11. doi: 10.1186/1471-2164-10-S1-S11. BMC Genomics. 2009. PMID: 19594870 Free PMC article.
-
Population Genomics Insights into Adaptive Evolution and Ecological Differentiation in Streptomycetes.Appl Environ Microbiol. 2019 Mar 22;85(7):e02555-18. doi: 10.1128/AEM.02555-18. Print 2019 Apr 1. Appl Environ Microbiol. 2019. PMID: 30658977 Free PMC article.
-
Evolutionary dynamics of rhomboid proteases in Streptomycetes.BMC Res Notes. 2015 Jun 9;8:234. doi: 10.1186/s13104-015-1205-x. BMC Res Notes. 2015. PMID: 26054641 Free PMC article.
-
Plasticity of the streptomyces genome-evolution and engineering of new antibiotics.Curr Med Chem. 2005;12(14):1697-704. doi: 10.2174/0929867054367176. Curr Med Chem. 2005. PMID: 16022666 Review.
-
Microbe Profile: Streptomyces coelicolor: a burlesque of pigments and phenotypes.Microbiology (Reading). 2019 Sep;165(9):953-955. doi: 10.1099/mic.0.000821. Epub 2019 Aug 1. Microbiology (Reading). 2019. PMID: 31368887 Free PMC article. Review.
Cited by
-
Evolution and Diversity of Assembly-Line Polyketide Synthases.Chem Rev. 2019 Dec 26;119(24):12524-12547. doi: 10.1021/acs.chemrev.9b00525. Epub 2019 Dec 15. Chem Rev. 2019. PMID: 31838842 Free PMC article. Review.
-
Accumulation of a bioactive benzoisochromanequinone compound kalafungin by a wild type antitumor-medermycin-producing streptomycete strain.PLoS One. 2015 Feb 19;10(2):e0117690. doi: 10.1371/journal.pone.0117690. eCollection 2015. PLoS One. 2015. PMID: 25695632 Free PMC article.
-
Deciphering the Transcriptional Response Mediated by the Redox-Sensing System HbpS-SenS-SenR from Streptomycetes.PLoS One. 2016 Aug 19;11(8):e0159873. doi: 10.1371/journal.pone.0159873. eCollection 2016. PLoS One. 2016. PMID: 27541358 Free PMC article.
-
Streptomyces coelicolor encodes a urate-responsive transcriptional regulator with homology to PecS from plant pathogens.J Bacteriol. 2013 Nov;195(21):4954-65. doi: 10.1128/JB.00854-13. Epub 2013 Aug 30. J Bacteriol. 2013. PMID: 23995633 Free PMC article.
-
Deciphering the streamlined genome of Streptomyces xiamenensis 318 as the producer of the anti-fibrotic drug candidate xiamenmycin.Sci Rep. 2016 Jan 8;6:18977. doi: 10.1038/srep18977. Sci Rep. 2016. PMID: 26744183 Free PMC article.
References
-
- Garrity GM, Lilburn TG, Cole JR, Harrison SH, Euzéby J, BJ T. Part 10 - The Bacteria: Phylum "Actinobacteria": Class Actinobacteria. Taxonomic Outline of the Bacteria and Archaea. 2007;Release 7.7.:399–539.
-
- Hopwood DA. Streptomyces in Nature and Medicine: The Antibiotic Makers. New York: Oxford University Press; 2007.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

