Differential function of the two Atg4 homologues in the aggrephagy pathway in Caenorhabditis elegans
- PMID: 22767594
- PMCID: PMC3436130
- DOI: 10.1074/jbc.M112.365676
Differential function of the two Atg4 homologues in the aggrephagy pathway in Caenorhabditis elegans
Abstract
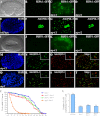
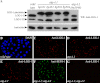
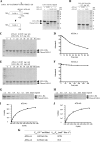
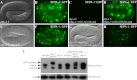
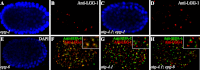
The presence of multiple homologues of the same yeast Atg protein endows an additional layer of complexity on the autophagy pathway in higher eukaryotes. The physiological function of the individual genes, however, remains largely unknown. Here we investigated the role of the two Caenorhabditis elegans homologues of the cysteine protease Atg4 in the pathway responsible for degradation of protein aggregates. Loss of atg-4.1 activity causes defective degradation of a variety of protein aggregates, whereas atg-4.2 mutants remove these substrates normally. LGG-1 precursors accumulate in atg-4.1 mutants, but not atg-4.2 mutants. LGG-1 puncta, formation of which depends on lipidation of LGG-1, are present in atg-4.1 and atg-4.2 single mutants, but are completely absent in atg-4.1; atg-4.2 double mutants. In vitro enzymatic analysis revealed that ATG-4.1 processes LGG-1 precursors about 100-fold more efficiently than ATG-4.2. Expression of a mutant form LGG-1, which mimics the processed precursor, rescues the defective autophagic degradation of protein aggregates in atg-4.1 mutants and, to a lesser extent, in atg-4.1; atg-4.2 double mutants. Our study reveals that ATG-4.1 and ATG-4.2 are functionally redundant yet display differential LGG-1 processing and deconjugating activity in the aggrephagy pathway in C. elegans.
Figures

References
-
- Xie Z., Klionsky D. J. (2007) Autophagosome formation: core machinery and adaptations. Nat. Cell Biol. 9, 1102–1109 - PubMed
-
- Nakatogawa H., Suzuki K., Kamada Y., Ohsumi Y. (2009) Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat. Rev. Mol. Cell Biol. 10, 458–467 - PubMed
-
- Ohsumi Y. (2001) Molecular dissection of autophagy: two ubiquitin-like systems. Nat. Rev. Mol. Cell Biol. 2, 211–216 - PubMed
-
- Kirisako T., Ichimura Y., Okada H., Kabeya Y., Mizushima N., Yoshimori T., Ohsumi M., Takao T., Noda T., Ohsumi Y. (2000) The reversible modification regulates the membrane-binding state of Apg8/Aut7 essential for autophagy and the cytoplasm to vacuole targeting pathway. J. Cell Biol. 151, 263–276 - PMC - PubMed
-
- Yoshimori T., Noda T. (2008) Toward unraveling membrane biogenesis in mammalian autophagy. Curr. Opin. Cell Biol. 20, 401–407 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

