Diversifying selection underlies the origin of allozyme polymorphism at the phosphoglucose isomerase locus in Tigriopus californicus
- PMID: 22768211
- PMCID: PMC3386920
- DOI: 10.1371/journal.pone.0040035
Diversifying selection underlies the origin of allozyme polymorphism at the phosphoglucose isomerase locus in Tigriopus californicus
Abstract
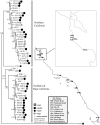
The marine copepod Tigriopus californicus lives in intertidal rock pools along the Pacific coast, where it exhibits strong, temporally stable population genetic structure. Previous allozyme surveys have found high frequency private alleles among neighboring subpopulations, indicating that there is limited genetic exchange between populations. Here we evaluate the factors responsible for the diversification and maintenance of alleles at the phosphoglucose isomerase (Pgi) locus by evaluating patterns of nucleotide variation underlying previously identified allozyme polymorphism. Copepods were sampled from eleven sites throughout California and Baja California, revealing deep genetic structure among populations as well as genetic variability within populations. Evidence of recombination is limited to the sample from Pescadero and there is no support for linkage disequilibrium across the Pgi locus. Neutrality tests and codon-based models of substitution suggest the action of natural selection due to elevated non-synonymous substitutions at a small number of sites in Pgi. Two sites are identified as the charge-changing residues underlying allozyme polymorphisms in T. californicus. A reanalysis of allozyme variation at several focal populations, spanning a period of 26 years and over 200 generations, shows that Pgi alleles are maintained without notable frequency changes. Our data suggest that diversifying selection accounted for the origin of Pgi allozymes, while McDonald-Kreitman tests and the temporal stability of private allozyme alleles suggests that balancing selection may be involved in the maintenance of amino acid polymorphisms within populations.
Conflict of interest statement
Figures
Similar articles
-
From DNA to fitness differences: sequences and structures of adaptive variants of Colias phosphoglucose isomerase (PGI).Mol Biol Evol. 2006 Mar;23(3):499-512. doi: 10.1093/molbev/msj062. Epub 2005 Nov 16. Mol Biol Evol. 2006. PMID: 16292000 Free PMC article.
-
Nucleotide polymorphism at a gene (Pgi) under balancing selection in a butterfly metapopulation.Mol Biol Evol. 2010 Feb;27(2):267-81. doi: 10.1093/molbev/msp227. Epub 2009 Sep 30. Mol Biol Evol. 2010. PMID: 19793833
-
Contrasting patterns of nucleotide polymorphism suggest different selective regimes within different parts of the PgiC1 gene in Festuca ovina L.Hereditas. 2017 May 18;154:11. doi: 10.1186/s41065-017-0032-6. eCollection 2017. Hereditas. 2017. PMID: 28529468 Free PMC article.
-
Genetic variation in the acorn barnacle from allozymes to population genomics.Integr Comp Biol. 2012 Sep;52(3):418-29. doi: 10.1093/icb/ics099. Epub 2012 Jul 4. Integr Comp Biol. 2012. PMID: 22767487 Free PMC article. Review.
-
Selection of allelic isozyme polymorphisms in marine organisms: pattern, theory, and application.Isozymes Curr Top Biol Med Res. 1983;10:69-92. Isozymes Curr Top Biol Med Res. 1983. PMID: 6226622 Review.
Cited by
-
Evidence for Positive Selection within the PgiC1 Locus in the Grass Festuca ovina.PLoS One. 2015 May 6;10(5):e0125831. doi: 10.1371/journal.pone.0125831. eCollection 2015. PLoS One. 2015. PMID: 25946223 Free PMC article.
-
Investigating the molecular basis of local adaptation to thermal stress: population differences in gene expression across the transcriptome of the copepod Tigriopus californicus.BMC Evol Biol. 2012 Sep 5;12:170. doi: 10.1186/1471-2148-12-170. BMC Evol Biol. 2012. PMID: 22950661 Free PMC article.
-
Fin whale MDH-1 and MPI allozyme variation is not reflected in the corresponding DNA sequences.Ecol Evol. 2014 May;4(10):1787-803. doi: 10.1002/ece3.1046. Epub 2014 Apr 16. Ecol Evol. 2014. PMID: 24963377 Free PMC article.
-
Positive selection in glycolysis among Australasian stick insects.BMC Evol Biol. 2013 Sep 30;13:215. doi: 10.1186/1471-2148-13-215. BMC Evol Biol. 2013. PMID: 24079656 Free PMC article.
-
Positive selection on human gamete-recognition genes.PeerJ. 2018 Jan 11;6:e4259. doi: 10.7717/peerj.4259. eCollection 2018. PeerJ. 2018. PMID: 29340252 Free PMC article.
References
-
- McKay JK, Latta RG. Adaptive population divergence: markers, QTL and traits. Trends Ecol Evol. 2002;17:285–291.
-
- Fraser D, Bernatchez L. Adaptive evolutionary conservation: towards a unified concept for defining conservation units. Mol Ecol. 2001;10:2741–2752. - PubMed
-
- Morin PA, Luikart G, Wayne RK the SNP workshop group. SNPs in ecology, evolution and conservation. Trends Ecol Evol. 2004;19:208–216.
-
- Ellegren H, Sheldon BC. Genetic basis of fitness differences in natural populations. Nature. 2008;452:169–175. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

