Schizosaccharomyces pombe Hat1 (Kat1) is associated with Mis16 and is required for telomeric silencing
- PMID: 22771823
- PMCID: PMC3445983
- DOI: 10.1128/EC.00123-12
Schizosaccharomyces pombe Hat1 (Kat1) is associated with Mis16 and is required for telomeric silencing
Abstract
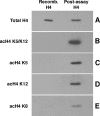
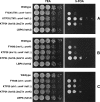
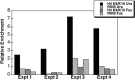
The Hat1 histone acetyltransferase has been implicated in the acetylation of histone H4 during chromatin assembly. In this study, we have characterized the Hat1 complex from the fission yeast Schizosaccharomyces pombe and have examined its role in telomeric silencing. Hat1 is found associated with the RbAp46 homologue Mis16, an essential protein. The Hat1 complex acetylates lysines 5 and 12 of histone H4, the sites that are acetylated in newly synthesized H4 in a wide range of eukaryotes. Deletion of hat1 in S. pombe is itself sufficient to cause the loss of silencing at telomeres. This is in contrast to results obtained with an S. cerevisiae hat1Δ strain, which must also carry mutations of specific acetylatable lysines in the H3 tail domain for loss of telomeric silencing to occur. Notably, deletion of hat1 from S. pombe resulted in an increase of acetylation of histone H4 in subtelomeric chromatin, concomitant with derepression of this region. A similar loss of telomeric silencing was also observed after growing cells in the presence of the deacetylase inhibitor trichostatin A. However, deleting hat1 did not cause loss of silencing at centromeres or the silent mating type locus. These results point to a direct link between Hat1, H4 acetylation, and the establishment of repressed telomeric chromatin in fission yeast.
Figures




References
-
- Ai X, Parthun MR. 2004. The nuclear Hat1p/Hat2p complex: a molecular link between type B histone acetyltransferases and chromatin assembly. Mol. Cell 14:195–205 - PubMed
-
- Allshire RC, Nimmo ER, Ekwall K, Javerzat JP, Cranston G. 1995. Mutations derepressing silent centromeric domains in fission yeast disrupt chromosome segregation. Genes Dev. 9:218–233 - PubMed
-
- Almeida R, Buscaino A, Allshire RC. 2006. Molecular biology: silencing unlimited. Curr. Biol. 16:R635–R638 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

