Poly(A) binding protein nuclear 1 levels affect alternative polyadenylation
- PMID: 22772983
- PMCID: PMC3467053
- DOI: 10.1093/nar/gks655
Poly(A) binding protein nuclear 1 levels affect alternative polyadenylation
Abstract
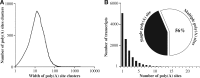
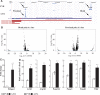
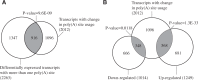
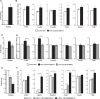
The choice for a polyadenylation site determines the length of the 3'-untranslated region (3'-UTRs) of an mRNA. Inclusion or exclusion of regulatory sequences in the 3'-UTR may ultimately affect gene expression levels. Poly(A) binding protein nuclear 1 (PABPN1) is involved in polyadenylation of pre-mRNAs. An alanine repeat expansion in PABPN1 (exp-PABPN1) causes oculopharyngeal muscular dystrophy (OPMD). We hypothesized that previously observed disturbed gene expression patterns in OPMD muscles may have been the result of an effect of PABPN1 on alternative polyadenylation, influencing mRNA stability, localization and translation. A single molecule polyadenylation site sequencing method was developed to explore polyadenylation site usage on a genome-wide level in mice overexpressing exp-PABPN1. We identified 2012 transcripts with altered polyadenylation site usage. In the far majority, more proximal alternative polyadenylation sites were used, resulting in shorter 3'-UTRs. 3'-UTR shortening was generally associated with increased expression. Similar changes in polyadenylation site usage were observed after knockdown or overexpression of expanded but not wild-type PABPN1 in cultured myogenic cells. Our data indicate that PABPN1 is important for polyadenylation site selection and that reduced availability of functional PABPN1 in OPMD muscles results in use of alternative polyadenylation sites, leading to large-scale deregulation of gene expression.
Figures


References
-
- Wahle E. Poly(A) tail length control is caused by termination of processive synthesis. J. Biol. Chem. 1995;270:2800–2808. - PubMed
-
- Benoit B, Mitou G, Chartier A, Temme C, Zaessinger S, Wahle E, Busseau I, Simonelig M. An essential cytoplasmic function for the nuclear poly(A) binding protein, PABP2, in poly(A) tail length control and early development in Drosophila. Dev. Cell. 2005;9:511–522. - PubMed
-
- Brais B, Bouchard JP, Xie YG, Rochefort DL, Chretien N, Tome FM, Lafreniere RG, Rommens JM, Uyama E, Nohira O, et al. Short GCG expansions in the PABP2 gene cause oculopharyngeal muscular dystrophy. Nat. Genet. 1998;18:164–167. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

