Mercury resistance and mercuric reductase activities and expression among chemotrophic thermophilic Aquificae
- PMID: 22773655
- PMCID: PMC3426723
- DOI: 10.1128/AEM.01060-12
Mercury resistance and mercuric reductase activities and expression among chemotrophic thermophilic Aquificae
Abstract
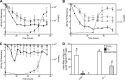
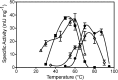
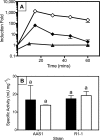
Mercury (Hg) resistance (mer) by the reduction of mercuric to elemental Hg is broadly distributed among the Bacteria and Archaea and plays an important role in Hg detoxification and biogeochemical cycling. MerA is the protein subunit of the homodimeric mercuric reductase (MR) enzyme, the central function of the mer system. MerA sequences in the phylum Aquificae form the deepest-branching lineage in Bayesian phylogenetic reconstructions of all known MerA homologs. We therefore hypothesized that the merA homologs in two thermophilic Aquificae, Hydrogenobaculum sp. strain Y04AAS1 (AAS1) and Hydrogenivirga sp. strain 128-5-R1-1 (R1-1), specified Hg resistance. Results supported this hypothesis, because strains AAS1 and R1-1 (i) were resistant to >10 μM Hg(II), (ii) transformed Hg(II) to Hg(0) during cellular growth, and (iii) possessed Hg-dependent NAD(P)H oxidation activities in crude cell extracts that were optimal at temperatures corresponding with the strains' optimal growth temperatures, 55°C for AAS1 and 70°C for R1-1. While these characteristics all conformed with the mer system paradigm, expression of the Aquificae mer operons was not induced by exposure to Hg(II) as indicated by unity ratios of merA transcripts, normalized to gyrA transcripts for hydrogen-grown AAS1 cultures, and by similar MR specific activities in thiosulfate-grown cultures with and without Hg(II). The Hg(II)-independent expression of mer in the deepest-branching lineage of MerA from bacteria whose natural habitats are Hg-rich geothermal environments suggests that regulated expression of mer was a later innovation likely in environments where microorganisms were intermittently exposed to toxic concentrations of Hg.
Figures

Similar articles
-
A thermophilic bacterial origin and subsequent constraints by redox, light and salinity on the evolution of the microbial mercuric reductase.Environ Microbiol. 2010 Nov;12(11):2904-17. doi: 10.1111/j.1462-2920.2010.02260.x. Environ Microbiol. 2010. PMID: 20545753
-
An initial characterization of the mercury resistance (mer) system of the thermophilic bacterium Thermus thermophilus HB27.FEMS Microbiol Ecol. 2009 Jan;67(1):118-29. doi: 10.1111/j.1574-6941.2008.00603.x. FEMS Microbiol Ecol. 2009. PMID: 19120462
-
Mercury adaptation among bacteria from a deep-sea hydrothermal vent.Appl Environ Microbiol. 2005 Jan;71(1):220-6. doi: 10.1128/AEM.71.1.220-226.2005. Appl Environ Microbiol. 2005. PMID: 15640191 Free PMC article.
-
Bacterial resistances to inorganic mercury salts and organomercurials.Plasmid. 1992 Jan;27(1):4-16. doi: 10.1016/0147-619x(92)90002-r. Plasmid. 1992. PMID: 1311113 Review.
-
Microbes in mercury-enriched geothermal springs in western North America.Sci Total Environ. 2016 Nov 1;569-570:321-331. doi: 10.1016/j.scitotenv.2016.06.080. Epub 2016 Jun 24. Sci Total Environ. 2016. PMID: 27344121 Review.
Cited by
-
Metalloid Reductase Activity Modified by a Fused Se0 Binding Peptide.ACS Chem Biol. 2020 Jul 17;15(7):1987-1995. doi: 10.1021/acschembio.0c00387. Epub 2020 Jul 6. ACS Chem Biol. 2020. PMID: 32568515 Free PMC article.
-
The mercury resistance operon: from an origin in a geothermal environment to an efficient detoxification machine.Front Microbiol. 2012 Oct 8;3:349. doi: 10.3389/fmicb.2012.00349. eCollection 2012. Front Microbiol. 2012. PMID: 23087676 Free PMC article.
-
Mercury-Tolerant Ensifer medicae Strains Display High Mercuric Reductase Activity and a Protective Effect on Nitrogen Fixation in Medicago truncatula Nodules Under Mercury Stress.Front Plant Sci. 2021 Jan 14;11:560768. doi: 10.3389/fpls.2020.560768. eCollection 2020. Front Plant Sci. 2021. PMID: 33519831 Free PMC article.
-
Comparative genomic analysis of phylogenetically closely related Hydrogenobaculum sp. isolates from Yellowstone National Park.Appl Environ Microbiol. 2013 May;79(9):2932-43. doi: 10.1128/AEM.03591-12. Epub 2013 Feb 22. Appl Environ Microbiol. 2013. PMID: 23435891 Free PMC article.
-
Molecular Adaptations of Bacterial Mercuric Reductase to the Hypersaline Kebrit Deep in the Red Sea.Appl Environ Microbiol. 2019 Feb 6;85(4):e01431-18. doi: 10.1128/AEM.01431-18. Print 2019 Feb 15. Appl Environ Microbiol. 2019. PMID: 30504211 Free PMC article.
References
-
- Andersson ME, Gardfeldt K, Wangberg I, Stromberg D. 2008. Determination of Henry's law constant for elemental mercury. Chemosphere 73:587–592 - PubMed
-
- Barkay T, Kritee K, Boyd E, Geesey GG. 2010. A thermophilic bacterial origin and subsequent constraints by redox, light and salinity on the evolution of the microbial mercuric reductase. Environ. Microbiol. 12:2904–2917 - PubMed
-
- Barkay T, Miller SM, Summers AO. 2003. Bacterial mercury resistance from atoms to ecosystems. FEMS Microbiol. Rev. 27:355–384 - PubMed
-
- Barkay T, Wagner-Döbler I. 2005. Microbial transformations of mercury: potentials, challenges, and achievements in controlling mercury toxicity in the environment. Adv. Appl. Microbiol. 57:1–52 - PubMed
-
- Ben-David EA, Holden PJ, Stone DJ, Harch BD, Foster LJ. 2004. The use of phospholipid fatty acid analysis to measure impact of acid rock drainage on microbial communities in sediments. Microb. Ecol. 48:300–315 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

