Defining desirable central nervous system drug space through the alignment of molecular properties, in vitro ADME, and safety attributes
- PMID: 22778836
- PMCID: PMC3368653
- DOI: 10.1021/cn100007x
Defining desirable central nervous system drug space through the alignment of molecular properties, in vitro ADME, and safety attributes
Abstract
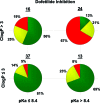
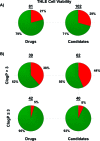
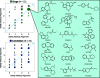
As part of our effort to increase survival of drug candidates and to move our medicinal chemistry design to higher probability space for success in the Neuroscience therapeutic area, we embarked on a detailed study of the property space for a collection of central nervous system (CNS) molecules. We carried out a thorough analysis of properties for 119 marketed CNS drugs and a set of 108 Pfizer CNS candidates. In particular, we focused on understanding the relationships between physicochemical properties, in vitro ADME (absorption, distribution, metabolism, and elimination) attributes, primary pharmacology binding efficiencies, and in vitro safety data for these two sets of compounds. This scholarship provides guidance for the design of CNS molecules in a property space with increased probability of success and may lead to the identification of druglike candidates with favorable safety profiles that can successfully test hypotheses in the clinic.
Keywords: CNS candidates; CNS drugs; Central nervous system (CNS); Madin−Darby canine kidney; P-glycoprotein; cellular toxicity; dofetilide binding; drug−drug interactions; high-throughput screening; human liver microsome stability; hydrogen bond donor; ligand efficiency; ligand-efficiency-dependent lipophilicity; ligand-lipophilicity efficiency; lipophilicity; molecular weight; most basic pKa; passive permeability; polarity; topological polar surface area; transformed human liver epithelial cells; unbound intrinsic clearance.
Figures

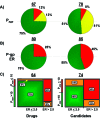

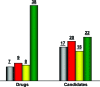

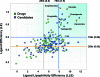
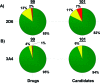




References
-
- Kola I.; Landis J. (2004) Opinion: Can the pharmaceutical industry reduce attrition rates?. Nat. Rev. Drug Discovery 3, 711–716. - PubMed
-
- O'Shea R.; Moser H. E. (2008) Physicochemical properties of antibacterial compounds: Implications for drug discovery. J. Med. Chem. 51, 2871–2878. - PubMed
-
- Leeson Paul D.; Davis Andrew M. (2004) Time-related differences in the physical property profiles of oral drugs. J. Med. Chem. 47, 6338–6348. - PubMed
-
- Veber D. F.; Johnson S. R.; Cheng H.-Y.; Smith B. R.; Ward K. W.; Kopple K. D. (2002) Molecular properties that influence the oral bioavailability of drug candidates. J. Med. Chem. 45, 2615–2623. - PubMed
-
- Proudfoot J. R. (2005) The evolution of synthetic oral drug properties. Bioorg. Med. Chem. Lett. 15, 1087–1090. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

