On the origin of the unusual behavior in the stretching of single-stranded DNA
- PMID: 22779622
- PMCID: PMC3391302
- DOI: 10.1063/1.4729371
On the origin of the unusual behavior in the stretching of single-stranded DNA
Abstract
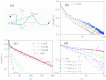
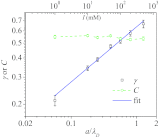
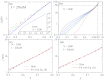
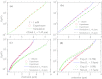
Force-extension curves (FECs), which quantify the response of a variety of biomolecules subject to mechanical force (f), are often quantitatively fit using worm-like chain (WLC) or freely jointed chain (FJC) models. These models predict that the chain extension, x, normalized by the contour length increases linearly at small f and at high forces scale as x ~ (1 - f(-α)), where α = 0.5 for WLC and unity for FJC. In contrast, experiments on single-stranded DNA (ssDNA) show that over a range of f and ionic concentration, x scales as x ~ ln f, which cannot be explained using WLC or FJC models. Using theory and simulations we show that this unusual behavior in FEC in ssDNA is due to sequence-independent polyelectrolyte effects. We show that the x ~ ln f arises because in the absence of force the tangent correlation function, quantifying chain persistence, decays algebraically on length scales on the order of the Debye length. Our theory, which is most appropriate for monovalent salts, quantitatively fits the experimental data and further predicts that such a regime is not discernible in double-stranded DNA.
Figures
Similar articles
-
Theory of Biopolymer Stretching at High Forces.Macromolecules. 2010 May 11;43(9):4394-4400. doi: 10.1021/ma902008y. Epub 2010 Apr 7. Macromolecules. 2010. PMID: 29225374 Free PMC article.
-
Mesoscopic models for DNA stretching under force: New results and comparison with experiments.Eur Phys J E Soft Matter. 2012 Oct;35(10):110. doi: 10.1140/epje/i2012-12110-2. Epub 2012 Oct 29. Eur Phys J E Soft Matter. 2012. PMID: 23099534
-
Elasticity of short DNA molecules: theory and experiment for contour lengths of 0.6-7 microm.Biophys J. 2007 Dec 15;93(12):4360-73. doi: 10.1529/biophysj.107.112995. Epub 2007 Aug 31. Biophys J. 2007. PMID: 17766363 Free PMC article.
-
Sugar-Pucker Force-Induced Transition in Single-Stranded DNA.Int J Mol Sci. 2021 Apr 29;22(9):4745. doi: 10.3390/ijms22094745. Int J Mol Sci. 2021. PMID: 33947069 Free PMC article.
-
Nanoscale mechanical and dynamical properties of DNA single molecules.Biophys Chem. 2005 Mar 1;113(3):209-21. doi: 10.1016/j.bpc.2004.09.007. Biophys Chem. 2005. PMID: 15620506 Review.
Cited by
-
Visualizing single-stranded nucleic acids in solution.Nucleic Acids Res. 2017 May 19;45(9):e66. doi: 10.1093/nar/gkw1297. Nucleic Acids Res. 2017. PMID: 28034955 Free PMC article.
-
Ab initio predictions for 3D structure and stability of single- and double-stranded DNAs in ion solutions.PLoS Comput Biol. 2022 Oct 19;18(10):e1010501. doi: 10.1371/journal.pcbi.1010501. eCollection 2022 Oct. PLoS Comput Biol. 2022. PMID: 36260618 Free PMC article.
-
The snakelike chain character of unstructured RNA.Biophys J. 2013 Dec 3;105(11):2569-76. doi: 10.1016/j.bpj.2013.10.019. Biophys J. 2013. PMID: 24314087 Free PMC article.
-
Global force-torque phase diagram for the DNA double helix: structural transitions, triple points, and collapsed plectonemes.Phys Rev E Stat Nonlin Soft Matter Phys. 2013 Dec;88(6):062722. doi: 10.1103/PhysRevE.88.062722. Epub 2013 Dec 27. Phys Rev E Stat Nonlin Soft Matter Phys. 2013. PMID: 24483501 Free PMC article.
-
Computational Modeling of DNA 3D Structures: From Dynamics and Mechanics to Folding.Molecules. 2023 Jun 17;28(12):4833. doi: 10.3390/molecules28124833. Molecules. 2023. PMID: 37375388 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

