A Lon-like protease with no ATP-powered unfolding activity
- PMID: 22792246
- PMCID: PMC3391209
- DOI: 10.1371/journal.pone.0040226
A Lon-like protease with no ATP-powered unfolding activity
Abstract
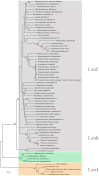
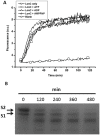
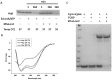
Lon proteases are a family of ATP-dependent proteases involved in protein quality control, with a unique proteolytic domain and an AAA(+) (ATPases associated with various cellular activities) module accommodated within a single polypeptide chain. They were classified into two types as either the ubiquitous soluble LonA or membrane-inserted archaeal LonB. In addition to the energy-dependent forms, a number of medically and ecologically important groups of bacteria encode a third type of Lon-like proteins in which the conserved proteolytic domain is fused to a large N-terminal fragment lacking canonical AAA(+) motifs. Here we showed that these Lon-like proteases formed a clade distinct from LonA and LonB. Characterization of one such Lon-like protease from Meiothermus taiwanensis indicated that it formed a hexameric assembly with a hollow chamber similar to LonA/B. The enzyme was devoid of ATPase activity but retained an ability to bind symmetrically six nucleotides per hexamer; accordingly, structure-based alignment suggested possible existence of a non-functional AAA-like domain. The enzyme degraded unstructured or unfolded protein and peptide substrates, but not well-folded proteins, in ATP-independent manner. These results highlight a new type of Lon proteases that may be involved in breakdown of excessive damage or unfolded proteins during stress conditions without consumption of energy.
Conflict of interest statement
Figures





Similar articles
-
Structural basis for the ATP-independent proteolytic activity of LonB proteases and reclassification of their AAA+ modules.J Microbiol. 2015 Oct;53(10):711-7. doi: 10.1007/s12275-015-5417-5. Epub 2015 Oct 2. J Microbiol. 2015. PMID: 26428922
-
Molecular insights into substrate recognition and discrimination by the N-terminal domain of Lon AAA+ protease.Elife. 2021 Apr 30;10:e64056. doi: 10.7554/eLife.64056. Elife. 2021. PMID: 33929321 Free PMC article.
-
The N-terminal substrate-recognition domain of a LonC protease exhibits structural and functional similarity to cytosolic chaperones.Acta Crystallogr D Biol Crystallogr. 2013 Sep;69(Pt 9):1789-97. doi: 10.1107/S090744491301500X. Epub 2013 Aug 17. Acta Crystallogr D Biol Crystallogr. 2013. PMID: 23999302
-
Slicing a protease: structural features of the ATP-dependent Lon proteases gleaned from investigations of isolated domains.Protein Sci. 2006 Aug;15(8):1815-28. doi: 10.1110/ps.052069306. Protein Sci. 2006. PMID: 16877706 Free PMC article. Review.
-
Structure and the Mode of Activity of Lon Proteases from Diverse Organisms.J Mol Biol. 2022 Apr 15;434(7):167504. doi: 10.1016/j.jmb.2022.167504. Epub 2022 Feb 17. J Mol Biol. 2022. PMID: 35183556 Free PMC article. Review.
Cited by
-
Structural basis for the ATP-independent proteolytic activity of LonB proteases and reclassification of their AAA+ modules.J Microbiol. 2015 Oct;53(10):711-7. doi: 10.1007/s12275-015-5417-5. Epub 2015 Oct 2. J Microbiol. 2015. PMID: 26428922
-
Processive cleavage of substrate at individual proteolytic active sites of the Lon protease complex.Sci Adv. 2021 Nov 12;7(46):eabj9537. doi: 10.1126/sciadv.abj9537. Epub 2021 Nov 10. Sci Adv. 2021. PMID: 34757797 Free PMC article.
-
Involvement of the N Domain Residues E34, K35, and R38 in the Functionally Active Structure of Escherichia coli Lon Protease.Acta Naturae. 2020 Oct-Dec;12(4):86-97. doi: 10.32607/actanaturae.11197. Acta Naturae. 2020. PMID: 33456980 Free PMC article.
-
Unique Structural Fold of LonBA Protease from Bacillus subtilis, a Member of a Newly Identified Subfamily of Lon Proteases.Int J Mol Sci. 2022 Sep 28;23(19):11425. doi: 10.3390/ijms231911425. Int J Mol Sci. 2022. PMID: 36232729 Free PMC article.
-
Modulation of Lonp1 Activity by Small Compounds.Biomolecules. 2025 Apr 9;15(4):553. doi: 10.3390/biom15040553. Biomolecules. 2025. PMID: 40305312 Free PMC article. Review.
References
-
- Sauer RT, Baker TA. AAA+ proteases: ATP-fueled machines of protein destruction. Ann. Review Biochem. 80, 587–612. 2011. - PubMed
-
- Neuwald AF, Aravind L, Spouge JL, Koonin EV. AAA+: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 9, 27–43. 1999. - PubMed
-
- Tsilibaris V, Maenhaut-Michel G, Van Melderen L. Biological roles of the Lon ATP-dependent protease. Res. Microbiol. 157, 701–713. 2006. - PubMed
-
- Goldberg AL, Moerschell RP, Chung CH, Maurizi MR. ATP-dependent protease La (lon) from Escherichia coli. Methods Enzymol. 244, 350–375. 1994. - PubMed
-
- Amerik A, Antonov VK, Gorbalenya AE, Kotova SA, Rotanova TV, et al. Site-directed mutagenesis of La protease. A catalytically active serine residue. FEBS. Lett. 287, 211–214. 1991. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

