Comparative genome analysis of the high pathogenicity Salmonella Typhimurium strain UK-1
- PMID: 22792393
- PMCID: PMC3391293
- DOI: 10.1371/journal.pone.0040645
Comparative genome analysis of the high pathogenicity Salmonella Typhimurium strain UK-1
Abstract
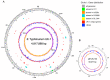
Salmonella enterica serovar Typhimurium, a gram-negative facultative rod-shaped bacterium causing salmonellosis and foodborne disease, is one of the most common isolated Salmonella serovars in both developed and developing nations. Several S. Typhimurium genomes have been completed and many more genome-sequencing projects are underway. Comparative genome analysis of the multiple strains leads to a better understanding of the evolution of S. Typhimurium and its pathogenesis. S. Typhimurium strain UK-1 (belongs to phage type 1) is highly virulent when orally administered to mice and chickens and efficiently colonizes lymphoid tissues of these species. These characteristics make this strain a good choice for use in vaccine development. In fact, UK-1 has been used as the parent strain for a number of nonrecombinant and recombinant vaccine strains, including several commercial vaccines for poultry. In this study, we conducted a thorough comparative genome analysis of the UK-1 strain with other S. Typhimurium strains and examined the phenotypic impact of several genomic differences. Whole genomic comparison highlights an extremely close relationship between the UK-1 strain and other S. Typhimurium strains; however, many interesting genetic and genomic variations specific to UK-1 were explored. In particular, the deletion of a UK-1-specific gene that is highly similar to the gene encoding the T3SS effector protein NleC exhibited a significant decrease in oral virulence in BALB/c mice. The complete genetic complements in UK-1, especially those elements that contribute to virulence or aid in determining the diversity within bacterial species, provide key information in evaluating the functional characterization of important genetic determinants and for development of vaccines.
Conflict of interest statement
Figures



Similar articles
-
Genomic Approaches for Understanding the Characteristics of Salmonella enterica subsp. enterica Serovar Typhimurium ST1120, Isolated from Swine Feces in Korea.J Microbiol Biotechnol. 2017 Nov 28;27(11):1983-1993. doi: 10.4014/jmb.1708.08027. J Microbiol Biotechnol. 2017. PMID: 29032651
-
Genomic Analysis of Salmonella enterica Serovar Typhimurium Characterizes Strain Diversity for Recent U.S. Salmonellosis Cases and Identifies Mutations Linked to Loss of Fitness under Nitrosative and Oxidative Stress.mBio. 2016 Mar 8;7(2):e00154. doi: 10.1128/mBio.00154-16. mBio. 2016. PMID: 26956590 Free PMC article.
-
Genomic characterization of endemic Salmonella enterica serovar Typhimurium and Salmonella enterica serovar I 4,[5],12:i:- isolated in Malaysia.Infect Genet Evol. 2018 Aug;62:109-121. doi: 10.1016/j.meegid.2018.04.027. Epub 2018 Apr 21. Infect Genet Evol. 2018. PMID: 29684710
-
Complete genome sequence of the universal killer Salmonella enterica Serovar Typhimurium UK-1 (ATCC 68169).J Bacteriol. 2011 Aug;193(15):4035-6. doi: 10.1128/JB.05224-11. Epub 2011 May 27. J Bacteriol. 2011. PMID: 21622747 Free PMC article.
-
So similar, yet so different: uncovering distinctive features in the genomes of Salmonella enterica serovars Typhimurium and Typhi.FEMS Microbiol Lett. 2010 Apr;305(1):1-13. doi: 10.1111/j.1574-6968.2010.01904.x. Epub 2010 Jan 20. FEMS Microbiol Lett. 2010. PMID: 20146749 Review.
Cited by
-
Outer membrane vesicles from flagellin-deficient Salmonella enterica serovar Typhimurium induce cross-reactive immunity and provide cross-protection against heterologous Salmonella challenge.Sci Rep. 2016 Oct 4;6:34776. doi: 10.1038/srep34776. Sci Rep. 2016. PMID: 27698383 Free PMC article.
-
Prediction of late-onset preeclampsia using plasma proteomics: a longitudinal multi-cohort study.Sci Rep. 2024 Dec 28;14(1):30813. doi: 10.1038/s41598-024-81277-2. Sci Rep. 2024. PMID: 39730472 Free PMC article.
-
A Salmonella type III effector, PipA, works in a different manner than the PipA family effectors GogA and GtgA.PLoS One. 2021 Mar 18;16(3):e0248975. doi: 10.1371/journal.pone.0248975. eCollection 2021. PLoS One. 2021. PMID: 33735297 Free PMC article.
-
A naturally occurring single nucleotide polymorphism in the Salmonella SPI-2 type III effector srfH/sseI controls early extraintestinal dissemination.PLoS One. 2012;7(9):e45245. doi: 10.1371/journal.pone.0045245. Epub 2012 Sep 18. PLoS One. 2012. PMID: 23028876 Free PMC article.
-
Distribution of Gifsy-3 and of variants of ST64B and Gifsy-1 prophages amongst Salmonella enterica Serovar Typhimurium isolates: evidence that combinations of prophages promote clonality.PLoS One. 2014 Jan 24;9(1):e86203. doi: 10.1371/journal.pone.0086203. eCollection 2014. PLoS One. 2014. PMID: 24475087 Free PMC article.
References
-
- CDC. Preliminary FoodNet data on the incidence of infection with pathogens transmitted commonly through food–10 states, 2009. MMWR Morb Mortal Wkly Rep. 2010;59:418–422. - PubMed
-
- Gordon MA, Banda HT, Gondwe M, Gordon SB, Boeree MJ, et al. Non-typhoidal salmonella bacteraemia among HIV-infected Malawian adults: high mortality and frequent recrudescence. AIDS. 2002;16:1633–1641. - PubMed
-
- Gordon MA, Graham SM, Walsh AL, Wilson L, Phiri A, et al. Epidemics of invasive Salmonella enterica serovar enteritidis and S. enterica Serovar typhimurium infection associated with multidrug resistance among adults and children in Malawi. Clin Infect Dis. 2008;46:963–969. - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

