An unbiased approach to identify genes involved in development in a turtle with temperature-dependent sex determination
- PMID: 22793670
- PMCID: PMC3434017
- DOI: 10.1186/1471-2164-13-308
An unbiased approach to identify genes involved in development in a turtle with temperature-dependent sex determination
Abstract
Background: Many reptiles exhibit temperature-dependent sex determination (TSD). The initial cue in TSD is incubation temperature, unlike genotypic sex determination (GSD) where it is determined by the presence of specific alleles (or genetic loci). We used patterns of gene expression to identify candidates for genes with a role in TSD and other developmental processes without making a priori assumptions about the identity of these genes (ortholog-based approach). We identified genes with sexually dimorphic mRNA accumulation during the temperature sensitive period of development in the Red-eared slider turtle (Trachemys scripta), a turtle with TSD. Genes with differential mRNA accumulation in response to estrogen (estradiol-17β; E(2)) exposure and developmental stages were also identified.
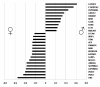
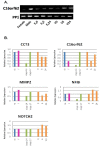
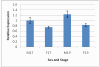
Results: Sequencing 767 clones from three suppression-subtractive hybridization libraries yielded a total of 581 unique sequences. Screening a macroarray with a subset of those sequences revealed a total of 26 genes that exhibited differential mRNA accumulation: 16 female biased and 10 male biased. Additional analyses revealed that C16ORF62 (an unknown gene) and MALAT1 (a long noncoding RNA) exhibited increased mRNA accumulation at the male producing temperature relative to the female producing temperature during embryonic sexual development. Finally, we identified four genes (C16ORF62, CCT3, MMP2, and NFIB) that exhibited a stage effect and five genes (C16ORF62, CCT3, MMP2, NFIB and NOTCH2) showed a response to E(2) exposure.
Conclusions: Here we report a survey of genes identified using patterns of mRNA accumulation during embryonic development in a turtle with TSD. Many previous studies have focused on examining the turtle orthologs of genes involved in mammalian development. Although valuable, the limitations of this approach are exemplified by our identification of two genes (MALAT1 and C16ORF62) that are sexually dimorphic during embryonic development. MALAT1 is a noncoding RNA that has not been implicated in sexual differentiation in other vertebrates and C16ORF62 has an unknown function. Our results revealed genes that are candidates for having roles in turtle embryonic development, including TSD, and highlight the need to expand our search parameters beyond protein-coding genes.
Figures

Similar articles
-
MeDIP-seq and nCpG analyses illuminate sexually dimorphic methylation of gonadal development genes with high historic methylation in turtle hatchlings with temperature-dependent sex determination.Epigenetics Chromatin. 2017 May 19;10:28. doi: 10.1186/s13072-017-0136-2. eCollection 2017. Epigenetics Chromatin. 2017. PMID: 28533820 Free PMC article.
-
Chronology, magnitude and duration of expression of putative sex-determining/differentiation genes in a turtle with temperature-dependent sex determination.Sex Dev. 2014;8(6):364-75. doi: 10.1159/000369116. Epub 2014 Nov 25. Sex Dev. 2014. PMID: 25427533
-
A timecourse analysis of systemic and gonadal effects of temperature on sexual development of the red-eared slider turtle Trachemys scripta elegans.Dev Biol. 2016 Dec 1;420(1):166-177. doi: 10.1016/j.ydbio.2016.09.018. Epub 2016 Sep 23. Dev Biol. 2016. PMID: 27671871
-
Temperature-dependent sex determination in the red-eared slider turtle, Trachemys scripta.J Exp Zool. 1998 Aug 1;281(5):409-16. doi: 10.1002/(sici)1097-010x(19980801)281:5<409::aid-jez6>3.0.co;2-s. J Exp Zool. 1998. PMID: 9662828 Review.
-
Steroid signaling and temperature-dependent sex determination-Reviewing the evidence for early action of estrogen during ovarian determination in turtles.Semin Cell Dev Biol. 2009 May;20(3):283-92. doi: 10.1016/j.semcdb.2008.10.004. Epub 2008 Nov 1. Semin Cell Dev Biol. 2009. PMID: 18992835 Free PMC article. Review.
Cited by
-
Survey of gene, lncRNA and transposon transcription patterns in four mouse organs highlights shared and organ-specific sex-biased regulation.Genome Biol. 2025 Mar 26;26(1):74. doi: 10.1186/s13059-025-03547-0. Genome Biol. 2025. PMID: 40140847 Free PMC article.
-
New locus reveals the genetic architecture of sex reversal in the Chinese tongue sole (Cynoglossus semilaevis).Heredity (Edinb). 2018 Oct;121(4):319-326. doi: 10.1038/s41437-018-0126-6. Epub 2018 Aug 9. Heredity (Edinb). 2018. PMID: 30093666 Free PMC article.
-
Single Locus Maintains Large Variation of Sex Reversal in Half-Smooth Tongue Sole (Cynoglossus semilaevis).G3 (Bethesda). 2017 Feb 9;7(2):583-589. doi: 10.1534/g3.116.036822. G3 (Bethesda). 2017. PMID: 28007836 Free PMC article.
-
Lasting effects of early exposure to temperature on the gonadal transcriptome at the time of sex differentiation in the European sea bass, a fish with mixed genetic and environmental sex determination.BMC Genomics. 2015 Sep 4;16(1):679. doi: 10.1186/s12864-015-1862-0. BMC Genomics. 2015. PMID: 26338702 Free PMC article.
-
Post-Transcriptional Mechanisms Respond Rapidly to Ecologically Relevant Thermal Fluctuations During Temperature-Dependent Sex Determination.Integr Org Biol. 2020 Oct 7;2(1):obaa033. doi: 10.1093/iob/obaa033. eCollection 2020. Integr Org Biol. 2020. PMID: 33791571 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

