The catalytic and the RNA subunits of human telomerase are required to immortalize equid primary fibroblasts
- PMID: 22797876
- PMCID: PMC3443485
- DOI: 10.1007/s00412-012-0379-4
The catalytic and the RNA subunits of human telomerase are required to immortalize equid primary fibroblasts
Abstract
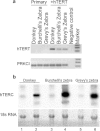
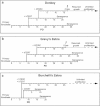
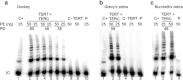
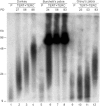
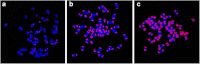
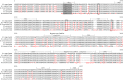
Many human primary somatic cells can be immortalized by inducing telomerase activity through the exogenous expression of the human telomerase catalytic subunit (hTERT). This approach has been extended to the immortalization of cell lines from several mammals. Here, we show that hTERT expression is not sufficient to immortalize primary fibroblasts from three equid species, namely donkey, Burchelli's zebra and Grevy's zebra. In vitro analysis of a reconstituted telomerase composed by hTERT and an equid RNA component of telomerase (TERC) revealed a low activity of this enzyme compared to human telomerase, suggesting a low compatibility of equid and human telomerase subunits. This conclusion was also strengthened by comparison of human and equid TERC sequences, which revealed nucleotide differences in key regions for TERC and TERT interaction. We then succeeded in immortalizing equid fibroblasts by expressing hTERT and hTERC concomitantly. Expression of both human telomerase subunits led to telomerase activity and telomere elongation, indicating that human telomerase is compatible with the other equid telomerase subunits and proteins involved in telomere metabolism. The immortalization procedure described herein could be extended to primary cells from other mammals. The availability of immortal cells from endangered species could be particularly useful for obtaining new information on the organization and function of their genomes, which is relevant for their preservation.
Figures



Similar articles
-
Ectopic expression of human telomerase RNA component results in increased telomerase activity and elongated telomeres in bovine blastocysts.Biol Reprod. 2012 Oct 25;87(4):95. doi: 10.1095/biolreprod.112.100198. Print 2012 Oct. Biol Reprod. 2012. PMID: 22855562
-
An alternate splicing variant of the human telomerase catalytic subunit inhibits telomerase activity.Neoplasia. 2000 Sep-Oct;2(5):433-40. doi: 10.1038/sj.neo.7900113. Neoplasia. 2000. PMID: 11191110 Free PMC article.
-
Mass cultured human fibroblasts overexpressing hTERT encounter a growth crisis following an extended period of proliferation.Exp Cell Res. 2000 Sep 15;259(2):336-50. doi: 10.1006/excr.2000.4982. Exp Cell Res. 2000. PMID: 10964501
-
Telomeres, telomerase, and myc. An update.Mutat Res. 2000 Jan;462(1):31-47. doi: 10.1016/s1383-5742(99)00091-5. Mutat Res. 2000. PMID: 10648922 Review.
-
Molecular regulation of telomerase activity in aging.Protein Cell. 2011 Sep;2(9):726-38. doi: 10.1007/s13238-011-1093-3. Epub 2011 Oct 6. Protein Cell. 2011. PMID: 21976062 Free PMC article. Review.
Cited by
-
Immortalization of American miniature horse-derived fibroblast by cell cycle regulator with normal karyotype.PeerJ. 2024 Jan 26;12:e16832. doi: 10.7717/peerj.16832. eCollection 2024. PeerJ. 2024. PMID: 38288466 Free PMC article.
-
Insertion of Telomeric Repeats in the Human and Horse Genomes: An Evolutionary Perspective.Int J Mol Sci. 2020 Apr 18;21(8):2838. doi: 10.3390/ijms21082838. Int J Mol Sci. 2020. PMID: 32325780 Free PMC article.
-
Expression of human mutant cyclin dependent kinase 4, Cyclin D and telomerase extends the life span but does not immortalize fibroblasts derived from loggerhead sea turtle (Caretta caretta).Sci Rep. 2018 Jun 20;8(1):9229. doi: 10.1038/s41598-018-27271-x. Sci Rep. 2018. PMID: 29925962 Free PMC article.
-
Molecular Dynamics and Evolution of Centromeres in the Genus Equus.Int J Mol Sci. 2022 Apr 10;23(8):4183. doi: 10.3390/ijms23084183. Int J Mol Sci. 2022. PMID: 35457002 Free PMC article. Review.
-
Birth, evolution, and transmission of satellite-free mammalian centromeric domains.Genome Res. 2018 Jun;28(6):789-799. doi: 10.1101/gr.231159.117. Epub 2018 Apr 30. Genome Res. 2018. PMID: 29712753 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

