Detecting cancer outlier genes with potential rearrangement using gene expression data and biological networks
- PMID: 22811706
- PMCID: PMC3394389
- DOI: 10.1155/2012/373506
Detecting cancer outlier genes with potential rearrangement using gene expression data and biological networks
Abstract
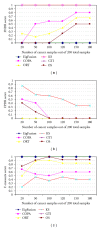
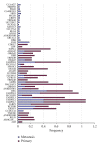
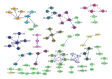
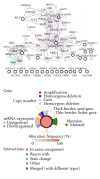
Gene alterations are a major component of the landscape of tumor genomes. To assess the significance of these alterations in the development of prostate cancer, it is necessary to identify these alterations and analyze them from systems biology perspective. Here, we present a new method (EigFusion) for predicting outlier genes with potential gene rearrangement. EigFusion demonstrated excellent performance in identifying outlier genes with potential rearrangement by testing it to synthetic and real data to evaluate performance. EigFusion was able to identify previously unrecognized genes such as FABP5 and KCNH8 and confirmed their association with primary and metastatic prostate samples while confirmed the metastatic specificity for other genes such as PAH, TOP2A, and SPINK1. We performed protein network based approaches to analyze the network context of potential rearranged genes. Functional gene rearrangement Modules are constructed by integrating functional protein networks. Rearranged genes showed to be highly connected to well-known altered genes in cancer such as AR, RB1, MYC, and BRCA1. Finally, using clinical outcome data of prostate cancer patients, potential rearranged genes demonstrated significant association with prostate cancer specific death.
Figures


References
-
- de Klein A, van Kessel AG, Grosveld G, et al. A celllular oncogene is translocated to the Philadelphia chromosome in chronic myelocytic leukaemia. Nature . 1982;300(5894):765–767. - PubMed
-
- Soda M, Choi YL, Enomoto M, et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature . 2007;448(7153):561–566. - PubMed
-
- Squire JA. TMPRSS2-ERG and PTEN loss in prostate cancer. Nature Genetics . 2009;41(5):509–510. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous

