FoxM1 regulates mammary luminal cell fate
- PMID: 22813746
- PMCID: PMC3401379
- DOI: 10.1016/j.celrep.2012.05.005
FoxM1 regulates mammary luminal cell fate
Abstract
Elevated expression of FoxM1 in breast cancer correlates with an undifferentiated tumor phenotype and a negative clinical outcome. However, a role for FoxM1 in regulating mammary differentiation was not known. Here, we identify another function of FoxM1, the ability to act as a transcriptional repressor, which plays an important role in regulating the differentiation of luminal epithelial progenitors. Regeneration of mammary glands with elevated levels of FoxM1 leads to aberrant ductal morphology and expansion of the luminal progenitor pool. Conversely, knockdown of FoxM1 results in a shift toward the differentiated state. FoxM1 mediates these effects by repressing the key regulator of luminal differentiation, GATA-3. Through association with DNMT3b, FoxM1 promotes methylation of the GATA-3 promoter in an Rb-dependent manner. This study identifies FoxM1 as a critical regulator of mammary differentiation with significant implications for the development of aggressive breast cancers.
Copyright © 2012 The Authors. Published by Elsevier Inc. All rights reserved.
Figures

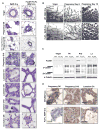





References
-
- Arnold A, Papanikolaou A. Cyclin D1 in breast cancer pathogenesis. J Clin Oncol. 2005;23:4215–24. - PubMed
-
- Asselin-Labat M, Sutherland K, Barker H, Thomas R, Shackleton M, Forrest N, Hartley L, Growveld F, Wees J, Lindeman G, Visvader J. Gata-3 is an essential regulator of mammary-gland morphogenesis and luminal-cell differentiation. Nat Cell Biol. 2007;9:201–9. - PubMed
-
- Asselin-Labat M, Vaillant F, Sheridan J, Pal B, Wu D, Simpson E, Yasuda H, Smyth G, Martin J, Lindeman GJ, Visvader J. Control of mammary stem cell function by steroid hormone signaling. Nature. 2010;465:798–802. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

