A global view of the oncogenic landscape in nasopharyngeal carcinoma: an integrated analysis at the genetic and expression levels
- PMID: 22815911
- PMCID: PMC3398876
- DOI: 10.1371/journal.pone.0041055
A global view of the oncogenic landscape in nasopharyngeal carcinoma: an integrated analysis at the genetic and expression levels
Abstract
Previous studies have reported that the tumour cells of nasopharyngeal carcinoma (NPC) exhibit recurrent chromosome abnormalities. These genetic changes are broadly assumed to lead to changes in gene expression which are important for the pathogenesis of this tumour. However, this assumption has yet to be formally tested at a global level. Therefore a genome wide analysis of chromosome copy number and gene expression was performed in tumour cells micro-dissected from the same NPC biopsies. Cellular tumour suppressor and tumour-promoting genes (TSG, TPG) and Epstein-Barr Virus (EBV)-encoded oncogenes were examined. The EBV-encoded genome maintenance protein EBNA1, along with the putative oncogenes LMP1, LMP2 and BARF1 were expressed in the majority of NPCs that were analysed. Significant downregulation of expression in an average of 76 cellular TSGs per tumour was found, whilst a per-tumour average of 88 significantly upregulated, TPGs occurred. The expression of around 60% of putative TPGs and TSGs was both up-and down-regulated in different types of cancer, suggesting that the simplistic classification of genes as TSGs or TPGs may not be entirely appropriate and that the concept of context-dependent onco-suppressors may be more extensive than previously recognised. No significant enrichment of TPGs within regions of frequent genomic gain was seen but TSGs were significantly enriched within regions of frequent genomic loss. It is suggested that loss of the FHIT gene may be a driver of NPC tumourigenesis. Notwithstanding the association of TSGs with regions of genomic loss, on a gene by gene basis and excepting homozygous deletions and high-level amplification, there is very little correlation between chromosomal copy number aberrations and expression levels of TSGs and TPGs in NPC.
Conflict of interest statement
Figures

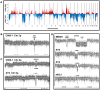

References
-
- Nicholls JM. Nasopharyngeal Carcinoma: Classification and histologic appearances. Advances in Anatomic Pathology. 1997;4:71–84.
-
- Chang ET, Adami HO. The enigmatic epidemiology of nasopharyngeal carcinoma. Cancer Epidemiol Biomarkers Prev. 2006;15:1765–1777. - PubMed
-
- Poirier S, Ohshima H, de-Thé G, Hubert A, Bourgade MC, et al. Volatile nitrosamine levels in common foods from Tunisia, south China and Greenland, high-risk areas for nasopharyngeal carcinoma (NPC). Int J Cancer. 1987;39:293–296. - PubMed
-
- Young LS, Rickinson AB. Epstein-Barr virus: 40 years on. Nature Rev. Cancer. 2004;4:757–768. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

