FAST-SeqS: a simple and efficient method for the detection of aneuploidy by massively parallel sequencing
- PMID: 22815955
- PMCID: PMC3399813
- DOI: 10.1371/journal.pone.0041162
FAST-SeqS: a simple and efficient method for the detection of aneuploidy by massively parallel sequencing
Abstract
Massively parallel sequencing of cell-free, maternal plasma DNA was recently demonstrated to be a safe and effective screening method for fetal chromosomal aneuploidies. Here, we report an improved sequencing method achieving significantly increased throughput and decreased cost by replacing laborious sequencing library preparation steps with PCR employing a single primer pair designed to amplify a discrete subset of repeated regions. Using this approach, samples containing as little as 4% trisomy 21 DNA could be readily distinguished from euploid samples.
Conflict of interest statement
Figures


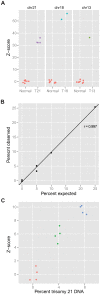
References
-
- Hsu L. Milunsky A, editor. Prenatal diagnosis of chromosomal abnormalities through amniocentesis. 1998. pp. 179–180. editor. Genetic Disorders and the Fetus. 4 ed. Baltimore: The Johns Hopkins University Press.
-
- Staebler M, Donner C, Van Regemorter N, Duprez L, De Maertelaer V, et al. Should determination of the karyotype be systematic for all malformations detected by obstetrical ultrasound? Prenat Diagn. 2005;25:567–573. - PubMed
-
- Jones KL. Smith’s recognizable patterns of human malformation. Philadelphia: Elsevier Saunders. 2006.
-
- American College of Obstetricians and Gynecologists. ACOG Practice Bulletin No. 88, December 2007. Invasive prenatal testing for aneuploidy. Obstet Gynecol. 2007;110:1459–1467. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials

