An α helix to β barrel domain switch transforms the transcription factor RfaH into a translation factor
- PMID: 22817892
- PMCID: PMC3430373
- DOI: 10.1016/j.cell.2012.05.042
An α helix to β barrel domain switch transforms the transcription factor RfaH into a translation factor
Abstract
NusG homologs regulate transcription and coupled processes in all living organisms. The Escherichia coli (E. coli) two-domain paralogs NusG and RfaH have conformationally identical N-terminal domains (NTDs) but dramatically different carboxy-terminal domains (CTDs), a β barrel in NusG and an α hairpin in RfaH. Both NTDs interact with elongating RNA polymerase (RNAP) to reduce pausing. In NusG, NTD and CTD are completely independent, and NusG-CTD interacts with termination factor Rho or ribosomal protein S10. In contrast, RfaH-CTD makes extensive contacts with RfaH-NTD to mask an RNAP-binding site therein. Upon RfaH interaction with its DNA target, the operon polarity suppressor (ops) DNA, RfaH-CTD is released, allowing RfaH-NTD to bind to RNAP. Here, we show that the released RfaH-CTD completely refolds from an all-α to an all-β conformation identical to that of NusG-CTD. As a consequence, RfaH-CTD binding to S10 is enabled and translation of RfaH-controlled operons is strongly potentiated. PAPERFLICK:
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures




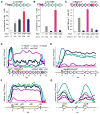


Comment in
-
Unfolding the bridge between transcription and translation.Cell. 2012 Jul 20;150(2):243-5. doi: 10.1016/j.cell.2012.06.025. Cell. 2012. PMID: 22817886 Free PMC article.
References
-
- Andreeva A, Murzin AG. Evolution of protein fold in the presence of functional constraints. Curr Opin Struct Biol. 2006;16:399–408. - PubMed
-
- Anfinsen CB. Principles that govern the folding of protein chains. Science. 1973;181:223–230. - PubMed
-
- Artsimovitch I, Landick R. The Transcriptional Regulator RfaH Stimulates RNA Chain Synthesis after Recruitment to Elongation Complexes by the Exposed Nontemplate DNA Strand. Cell. 2002;109:193–203. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

