Epigenetic genes regulated by the BRAFV600E signaling are associated with alterations in the methylation and expression of tumor suppressor genes and patient survival in melanoma
- PMID: 22820187
- PMCID: PMC3454505
- DOI: 10.1016/j.bbrc.2012.07.046
Epigenetic genes regulated by the BRAFV600E signaling are associated with alterations in the methylation and expression of tumor suppressor genes and patient survival in melanoma
Abstract
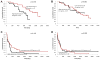
We have previously reported that the BRAFV600E signaling causes genome-wide aberrations in gene methylation in melanoma cells. To explore the potential molecular mechanisms for this epigenetic effect of BRAFV600E, in this in silico study we analyzed 11 microarray datasets retrieved from NCBI GEO database and examined the relationship of the expression of the epigenetic genes (genes involved in epigenetic regulation) with BRAFV600E signaling, methylation and expression of tumor-suppressor genes (TSGs) in melanoma, and patient survival with this cancer. Among 273 epigenetic genes examined, 12 genes were down-regulated (named DD genes) and 16 were up-regulated (UU genes) by suppression of the BRAFV600E signaling using inhibitors. While the expression of 245 non-DD/UU genes overall had no correlation with the expression and methylation of a set of potential TSGs, the expression of DD genes was significantly correlated negatively with the TSG expression and positively with TSG methylation. Expression of UU genes was positively, albeit weakly, associated with the TSG expression. Overall, no correlation was found between UU gene expression and TSG methylation. Importantly, the expression of DD genes, but not UU genes, was significantly associated with decreased survival of patients with melanoma. Interestingly, the promoters of DD genes contain more binding motifs of c-fos and myc, two BRAFV600E signaling-related transcription factors, than those of UU and non-DD/UU genes. Thus, these results link epigenetic genes to methylation and suppression of tumor suppressor genes as a mechanism involved in BRAFV600E-promoted melanoma tumorigenesis and uncover a novel molecular signature that predicts a poor prognosis of melanoma.
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures



Similar articles
-
The BRAF(V600E) causes widespread alterations in gene methylation in the genome of melanoma cells.Cell Cycle. 2012 Jan 15;11(2):286-95. doi: 10.4161/cc.11.2.18707. Epub 2012 Jan 15. Cell Cycle. 2012. PMID: 22189819 Free PMC article.
-
In-depth genomic data analyses revealed complex transcriptional and epigenetic dysregulations of BRAFV600E in melanoma.Mol Cancer. 2015 Mar 14;14:60. doi: 10.1186/s12943-015-0328-y. Mol Cancer. 2015. PMID: 25890285 Free PMC article.
-
Epigenetic inactivation of tumor suppressor genes in serum of patients with cutaneous melanoma.J Invest Dermatol. 2006 Feb;126(2):422-31. doi: 10.1038/sj.jid.5700073. J Invest Dermatol. 2006. PMID: 16374457
-
Epigenetic regulation in human melanoma: past and future.Epigenetics. 2015;10(2):103-21. doi: 10.1080/15592294.2014.1003746. Epigenetics. 2015. PMID: 25587943 Free PMC article. Review.
-
Bona Fide Tumor Suppressor Genes Hypermethylated in Melanoma: A Narrative Review.Int J Mol Sci. 2021 Oct 1;22(19):10674. doi: 10.3390/ijms221910674. Int J Mol Sci. 2021. PMID: 34639015 Free PMC article. Review.
Cited by
-
Activities of multiple cancer-related pathways are associated with BRAF mutation and predict the resistance to BRAF/MEK inhibitors in melanoma cells.Cell Cycle. 2014;13(2):208-19. doi: 10.4161/cc.26971. Epub 2013 Oct 29. Cell Cycle. 2014. PMID: 24200969 Free PMC article.
-
DNA methylation characteristics of primary melanomas with distinct biological behaviour.PLoS One. 2014 May 15;9(5):e96612. doi: 10.1371/journal.pone.0096612. eCollection 2014. PLoS One. 2014. PMID: 24832207 Free PMC article.
-
Clinically relevant genes and regulatory pathways associated with NRASQ61 mutations in melanoma through an integrative genomics approach.Oncotarget. 2015 Feb 10;6(4):2496-508. doi: 10.18632/oncotarget.2954. Oncotarget. 2015. PMID: 25537510 Free PMC article.
-
REC8 is a novel tumor suppressor gene epigenetically robustly targeted by the PI3K pathway in thyroid cancer.Oncotarget. 2015 Nov 17;6(36):39211-24. doi: 10.18632/oncotarget.5391. Oncotarget. 2015. PMID: 26472282 Free PMC article.
-
Mutations in KMT2C, BCOR and KDM5C Predict Response to Immune Checkpoint Blockade Therapy in Non-Small Cell Lung Cancer.Cancers (Basel). 2022 Jun 6;14(11):2816. doi: 10.3390/cancers14112816. Cancers (Basel). 2022. PMID: 35681795 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

