Evidence for cyclic Di-GMP-mediated signaling in Bacillus subtilis
- PMID: 22821967
- PMCID: PMC3430322
- DOI: 10.1128/JB.01092-12
Evidence for cyclic Di-GMP-mediated signaling in Bacillus subtilis
Abstract
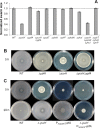
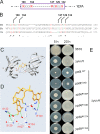
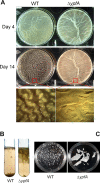
Cyclic di-GMP (c-di-GMP) is a second messenger that regulates diverse cellular processes in bacteria, including motility, biofilm formation, cell-cell signaling, and host colonization. Studies of c-di-GMP signaling have chiefly focused on Gram-negative bacteria. Here, we investigated c-di-GMP signaling in the Gram-positive bacterium Bacillus subtilis by constructing deletion mutations in genes predicted to be involved in the synthesis, breakdown, or response to the second messenger. We found that a putative c-di-GMP-degrading phosphodiesterase, YuxH, and a putative c-di-GMP receptor, YpfA, had strong influences on motility and that these effects depended on sequences similar to canonical EAL and RxxxR-D/NxSxxG motifs, respectively. Evidence indicates that YpfA inhibits motility by interacting with the flagellar motor protein MotA and that yuxH is under the negative control of the master regulator Spo0A∼P. Based on these findings, we propose that YpfA inhibits motility in response to rising levels of c-di-GMP during entry into stationary phase due to the downregulation of yuxH by Spo0A∼P. We also present evidence that YpfA has a mild influence on biofilm formation. In toto, our results demonstrate the existence of a functional c-di-GMP signaling system in B. subtilis that directly inhibits motility and directly or indirectly influences biofilm formation.
Figures





References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

