VirB-mediated positive feedback control of the virulence gene regulatory cascade of Shigella flexneri
- PMID: 22821978
- PMCID: PMC3457218
- DOI: 10.1128/JB.00800-12
VirB-mediated positive feedback control of the virulence gene regulatory cascade of Shigella flexneri
Abstract
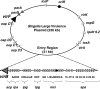
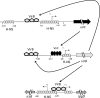
Shigella flexneri is a facultative intracellular pathogen that relies on a type III secretion system and its associated effector proteins to cause bacillary dysentery in humans. The genes that encode this virulence system are located on a 230-kbp plasmid and are transcribed in response to thermal, osmotic, and pH signals that are characteristic of the human lower gut. The virulence genes are organized within a regulatory cascade, and the nucleoid-associated protein H-NS represses each of the key promoters. Transcription derepression depends first on the VirF AraC-like transcription factor, a protein that antagonizes H-NS-mediated repression at the intermediate regulatory gene virB. The VirB protein in turn remodels the H-NS-DNA nucleoprotein complexes at the promoters of the genes encoding the type III secretion system and effector proteins, causing these genes to become derepressed. In this study, we show that the VirB protein also positively regulates the expression of its own gene (virB) via a cis-acting regulatory sequence. In addition, VirB positively regulates the gene coding for the VirF protein. This study reveals two hitherto uncharacterized feedback regulatory loops in the S. flexneri virulence cascade that provide a mechanism for the enhanced expression of the principal virulence regulatory genes.
Figures


References
-
- Adler B, et al. 1989. A dual transcriptional activation system for the 230-kb plasmid genes coding for virulence-associated antigens of Shigella flexneri. Mol. Microbiol. 3:627–635 - PubMed
-
- Allaoui A, Sansonetti PJ, Parsot C. 1993. MxiD, an outer membrane protein necessary for the secretion of the Shigella flexneri Ipa invasins. Mol. Microbiol. 7:59–68 - PubMed
-
- Beloin C, McKenna S, Dorman CJ. 2002. Molecular dissection of VirB, a key regulator of the virulence cascade of Shigella flexneri. J. Biol. Chem. 277:15333–15344 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

