Crystal structure of Δ-[Ru(bpy)₂dppz]²⁺ bound to mismatched DNA reveals side-by-side metalloinsertion and intercalation
- PMID: 22824892
- PMCID: PMC3478149
- DOI: 10.1038/nchem.1375
Crystal structure of Δ-[Ru(bpy)₂dppz]²⁺ bound to mismatched DNA reveals side-by-side metalloinsertion and intercalation
Abstract
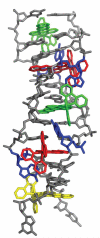
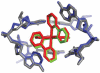
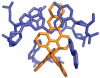
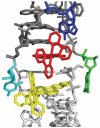
DNA mismatches represent a novel target in the development of diagnostics and therapeutics for cancer, because deficiencies in DNA mismatch repair are implicated in cancers, and cells that are repair-deficient show a high frequency of mismatches. Metal complexes with bulky intercalating ligands serve as probes for DNA mismatches. Here, we report the high-resolution (0.92 Å) crystal structure of the ruthenium 'light switch' complex Δ-[Ru(bpy)(2)dppz](2+) (bpy = 2,2'-bipyridine and dppz = dipyridophenazine), which is known to show luminescence on binding to duplex DNA, bound to both mismatched and well-matched sites in the oligonucleotide 5'-(dCGGAAATTACCG)(2)-3' (underline denotes AA mismatches). Two crystallographically independent views reveal that the complex binds mismatches through metalloinsertion, ejecting both mispaired adenosines. Additional ruthenium complexes are intercalated at well-matched sites, creating an array of complexes in the minor groove stabilized by stacking interactions between bpy ligands and extruded adenosines. This structure attests to the generality of metalloinsertion and metallointercalation as DNA binding modes.
Figures



Comment in
-
Interactions with DNA: Into the minor groove.Nat Chem. 2012 Jul 24;4(8):594-5. doi: 10.1038/nchem.1413. Nat Chem. 2012. PMID: 22824886 No abstract available.
References
-
- Loeb LA. A mutator phenotype in cancer. Cancer Res. 2001;61:3230–3239. - PubMed
-
- Strauss BS. Frameshift mutation, microsatellites and mismatch repair. Mutat. Res. 1999;437:195–203. - PubMed
-
- Papadopoulos N, Lindblom A. Molecular basis of HNPCC: mutations of MMR genes. Hum. Mutat. 1997;10:89–99. - PubMed
-
- Peltomaki P. Deficient DNA mismatch repair: a common etiologic factor for colon cancer. Hum. Mol. Genet. 2001;10:735–740. - PubMed
-
- Lawes DA, SenGupta S, Boulos PB. The clinical importance and prognostic implications of microsatellite instability in sporadic cancer. Eur. J. Surg. Oncol. 2003;29:201–212. - PubMed
Publication types
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

