Optimization of gene expression through divergent mutational paths
- PMID: 22832162
- PMCID: PMC3407975
- DOI: 10.1016/j.celrep.2011.12.003
Optimization of gene expression through divergent mutational paths
Abstract
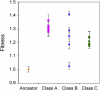
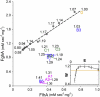
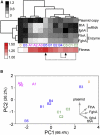
Adaptation under similar selective pressure often leads to comparable phenotypes. A longstanding question is whether such phenotypic repeatability entails similar (parallelism) or different genotypic changes (convergence). To better understand this, we characterized mutations that optimized expression of a plasmid-borne metabolic pathway during laboratory evolution of a bacterium. Expressing these pathway genes was essential for growth but came with substantial costs. Starting from overexpression, replicate populations founded by this bacterium all evolved to reduce expression. Despite this phenotypic repetitiveness, the underlying mutational spectrum was highly diverse. Analysis of these plasmid mutations identified three distinct means to modulate gene expression: (1) reducing the gene copy number, (2) lowering transcript stability, and (3) integration of the pathway-bearing plasmid into the host genome. Our study revealed diverse molecular changes beneath convergence to a simple phenotype. This complex genotype-phenotype mapping presents a challenge to inferring genetic evolution based solely on phenotypic changes.
Copyright © 2012 The Authors. Published by Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Microbial phenotypic heterogeneity in response to a metabolic toxin: Continuous, dynamically shifting distribution of formaldehyde tolerance in Methylobacterium extorquens populations.PLoS Genet. 2019 Nov 11;15(11):e1008458. doi: 10.1371/journal.pgen.1008458. eCollection 2019 Nov. PLoS Genet. 2019. PMID: 31710603 Free PMC article.
-
Fast growth increases the selective advantage of a mutation arising recurrently during evolution under metal limitation.PLoS Genet. 2009 Sep;5(9):e1000652. doi: 10.1371/journal.pgen.1000652. Epub 2009 Sep 18. PLoS Genet. 2009. PMID: 19763169 Free PMC article.
-
Diminishing returns epistasis among beneficial mutations decelerates adaptation.Science. 2011 Jun 3;332(6034):1190-2. doi: 10.1126/science.1203799. Science. 2011. PMID: 21636771 Free PMC article.
-
Breeding of Methanol-Tolerant Methylobacterium extorquens AM1 by Atmospheric and Room Temperature Plasma Mutagenesis Combined With Adaptive Laboratory Evolution.Biotechnol J. 2018 Jun;13(6):e1700679. doi: 10.1002/biot.201700679. Epub 2018 May 17. Biotechnol J. 2018. PMID: 29729127
-
Methylobacterium extorquens: methylotrophy and biotechnological applications.Appl Microbiol Biotechnol. 2015 Jan;99(2):517-34. doi: 10.1007/s00253-014-6240-3. Epub 2014 Nov 30. Appl Microbiol Biotechnol. 2015. PMID: 25432674 Review.
Cited by
-
It Takes Two to Make a Thing Go Right: Epistasis, Two-Component Response Systems, and Bacterial Adaptation.Microorganisms. 2024 Sep 30;12(10):2000. doi: 10.3390/microorganisms12102000. Microorganisms. 2024. PMID: 39458309 Free PMC article.
-
Methenyl-Dephosphotetrahydromethanopterin Is a Regulatory Signal for Acclimation to Changes in Substrate Availability in Methylobacterium extorquens AM1.J Bacteriol. 2015 Jun 15;197(12):2020-6. doi: 10.1128/JB.02595-14. Epub 2015 Apr 6. J Bacteriol. 2015. PMID: 25845846 Free PMC article.
-
Evolutionary adaptation after crippling cell polarization follows reproducible trajectories.Elife. 2015 Oct 1;4:e09638. doi: 10.7554/eLife.09638. Elife. 2015. PMID: 26426479 Free PMC article.
-
Good codons, bad transcript: large reductions in gene expression and fitness arising from synonymous mutations in a key enzyme.Mol Biol Evol. 2013 Mar;30(3):549-60. doi: 10.1093/molbev/mss273. Epub 2012 Dec 4. Mol Biol Evol. 2013. PMID: 23223712 Free PMC article.
-
Phenotypic and genotypic convergences are influenced by historical contingency and environment in yeast.Evolution. 2014 Mar;68(3):772-790. doi: 10.1111/evo.12302. Epub 2013 Nov 25. Evolution. 2014. PMID: 24164389 Free PMC article.
References
-
- Abzhanov A, Kuo WP, Hartmann C, Grant BR, Grant PR, Tabin CJ. The calmodulin pathway and evolution of elongated beak morphology in Darwin's finches. Nature. 2006;442:563–567. - PubMed
-
- Andersson DI, Hughes D. Gene amplification and adaptive evolution in bacteria. Annu. Rev. Genet. 2009;43:167–195. - PubMed
-
- Barrick JE, Yu DS, Yoon SH, Jeong H, Oh TK, Schneider D, Lenski RE, Kim JF. Genome evolution and adaptation in a long-term experiment with Escherichia coli. Nature. 2009;461:1243–1247. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

