Subcellular knockout of importin β1 perturbs axonal retrograde signaling
- PMID: 22841314
- PMCID: PMC3408616
- DOI: 10.1016/j.neuron.2012.05.033
Subcellular knockout of importin β1 perturbs axonal retrograde signaling
Abstract
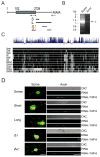
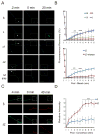
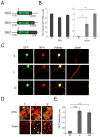
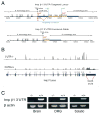
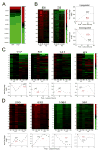
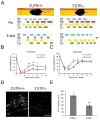
Subcellular localization of mRNA enables compartmentalized regulation within large cells. Neurons are the longest known cells; however, so far, evidence is lacking for an essential role of endogenous mRNA localization in axons. Localized upregulation of Importin β1 in lesioned axons coordinates a retrograde injury-signaling complex transported to the neuronal cell body. Here we show that a long 3' untranslated region (3' UTR) directs axonal localization of Importin β1. Conditional targeting of this 3' UTR region in mice causes subcellular loss of Importin β1 mRNA and protein in axons, without affecting cell body levels or nuclear functions in sensory neurons. Strikingly, axonal knockout of Importin β1 attenuates cell body transcriptional responses to nerve injury and delays functional recovery in vivo. Thus, localized translation of Importin β1 mRNA enables separation of cytoplasmic and nuclear transport functions of importins and is required for efficient retrograde signaling in injured axons.
Copyright © 2012 Elsevier Inc. All rights reserved.
Figures

Similar articles
-
Axonal transcription factors signal retrogradely in lesioned peripheral nerve.EMBO J. 2012 Mar 21;31(6):1350-63. doi: 10.1038/emboj.2011.494. Epub 2012 Jan 13. EMBO J. 2012. PMID: 22246183 Free PMC article.
-
Locally translated mTOR controls axonal local translation in nerve injury.Science. 2018 Mar 23;359(6382):1416-1421. doi: 10.1126/science.aan1053. Science. 2018. PMID: 29567716 Free PMC article.
-
Direct observation of importin α family member KPNA1 in axonal transport with or without a schizophrenia-related mutation.J Biol Chem. 2025 Apr;301(4):108343. doi: 10.1016/j.jbc.2025.108343. Epub 2025 Feb 24. J Biol Chem. 2025. PMID: 40010609 Free PMC article.
-
Integration of retrograde axonal and nuclear transport mechanisms in neurons: implications for therapeutics.Neuroscientist. 2004 Oct;10(5):404-8. doi: 10.1177/1073858404267884. Neuroscientist. 2004. PMID: 15359007 Review.
-
Local translation in neuronal processes--in vivo tests of a "heretical hypothesis".Dev Neurobiol. 2014 Mar;74(3):210-7. doi: 10.1002/dneu.22115. Epub 2013 Oct 11. Dev Neurobiol. 2014. PMID: 23929725 Review.
Cited by
-
Epac1 interacts with importin β1 and controls neurite outgrowth independently of cAMP and Rap1.Sci Rep. 2016 Nov 3;6:36370. doi: 10.1038/srep36370. Sci Rep. 2016. PMID: 27808165 Free PMC article.
-
Injury-induced HDAC5 nuclear export is essential for axon regeneration.Cell. 2013 Nov 7;155(4):894-908. doi: 10.1016/j.cell.2013.10.004. Cell. 2013. PMID: 24209626 Free PMC article.
-
Intra-axonal mechanisms driving axon regeneration.Brain Res. 2020 Aug 1;1740:146864. doi: 10.1016/j.brainres.2020.146864. Epub 2020 Apr 28. Brain Res. 2020. PMID: 32360100 Free PMC article. Review.
-
Advances in peripheral nerve regeneration.Nat Rev Neurol. 2013 Dec;9(12):668-76. doi: 10.1038/nrneurol.2013.227. Epub 2013 Nov 12. Nat Rev Neurol. 2013. PMID: 24217518 Review.
-
Deficiency in KPNA4, but Not in KPNA3, Causes Attention Deficit/Hyperactivity Disorder like Symptoms in Mice.Genes (Basel). 2025 Jun 6;16(6):690. doi: 10.3390/genes16060690. Genes (Basel). 2025. PMID: 40565582 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials

