Targeting a pre-mRNA structure with bipartite antisense molecules modulates tau alternative splicing
- PMID: 22844088
- PMCID: PMC3479178
- DOI: 10.1093/nar/gks710
Targeting a pre-mRNA structure with bipartite antisense molecules modulates tau alternative splicing
Abstract
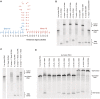
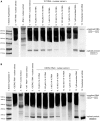
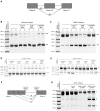
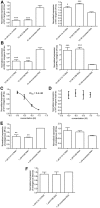
Approximately 15% of human genetic diseases are estimated to involve dysregulation of alternative pre-mRNA splicing. Antisense molecules designed to alter these and other splicing events typically target continuous linear sequences of the message. Here, we show that a structural feature in a pre-mRNA can be targeted by bipartite antisense molecules designed to hybridize with the discontinuous elements that flank the structure and thereby alter splicing. We targeted a hairpin structure at the boundary between exon 10 and intron 10 of the pre-mRNA of tau. Mutations in this region that are associated with certain forms of frontotemporal dementia, destabilize the hairpin to cause increased inclusion of exon 10. Via electrophoretic mobility shift and RNase protection assays, we demonstrate that bipartite antisense molecules designed to simultaneously interact with the available sequences that immediately flank the tau pre-mRNA hairpin do indeed bind to this structured region. Moreover, these agents inhibit exon 10 splicing and reverse the effect of destabilizing disease-causing mutations, in both in vitro splicing assays and cell culture. This general bipartite antisense strategy could be employed to modulate other splicing events that are regulated by RNA secondary structure.
Figures


Similar articles
-
Structural determinants for alternative splicing regulation of the MAPT pre-mRNA.RNA Biol. 2015;12(3):330-42. doi: 10.1080/15476286.2015.1017214. RNA Biol. 2015. PMID: 25826665 Free PMC article.
-
RNA Secondary Structure-Based Design of Antisense Peptide Nucleic Acids for Modulating Disease-Associated Aberrant Tau Pre-mRNA Alternative Splicing.Molecules. 2019 Aug 20;24(16):3020. doi: 10.3390/molecules24163020. Molecules. 2019. PMID: 31434312 Free PMC article.
-
Template-directed synthesis of a small molecule-antisense conjugate targeting an mRNA structure.Bioorg Chem. 2014 Jun;54:7-11. doi: 10.1016/j.bioorg.2014.03.001. Epub 2014 Mar 13. Bioorg Chem. 2014. PMID: 24691171 Free PMC article.
-
More than a messenger: Alternative splicing as a therapeutic target.Biochim Biophys Acta Gene Regul Mech. 2019 Nov-Dec;1862(11-12):194395. doi: 10.1016/j.bbagrm.2019.06.006. Epub 2019 Jul 2. Biochim Biophys Acta Gene Regul Mech. 2019. PMID: 31271898 Free PMC article. Review.
-
Splicing in the immune system: potential targets for therapeutic intervention by antisense-mediated alternative splicing.Curr Opin Mol Ther. 2009 Apr;11(2):124-32. Curr Opin Mol Ther. 2009. PMID: 19330718 Review.
Cited by
-
The Druggable Transcriptome Project: From Chemical Probes to Precision Medicines.Biochemistry. 2025 Apr 15;64(8):1647-1661. doi: 10.1021/acs.biochem.5c00006. Epub 2025 Mar 25. Biochemistry. 2025. PMID: 40131857 Review.
-
Splice-switching antisense oligonucleotides as therapeutic drugs.Nucleic Acids Res. 2016 Aug 19;44(14):6549-63. doi: 10.1093/nar/gkw533. Epub 2016 Jun 10. Nucleic Acids Res. 2016. PMID: 27288447 Free PMC article. Review.
-
RNA-based therapeutics for neurological diseases.RNA Biol. 2022;19(1):176-190. doi: 10.1080/15476286.2021.2021650. Epub 2021 Dec 31. RNA Biol. 2022. PMID: 35067193 Free PMC article. Review.
-
An intronic RNA element modulates Factor VIII exon-16 splicing.Nucleic Acids Res. 2024 Jan 11;52(1):300-315. doi: 10.1093/nar/gkad1034. Nucleic Acids Res. 2024. PMID: 37962303 Free PMC article.
-
Design, Optimization, and Study of Small Molecules That Target Tau Pre-mRNA and Affect Splicing.J Am Chem Soc. 2020 May 13;142(19):8706-8727. doi: 10.1021/jacs.0c00768. Epub 2020 May 4. J Am Chem Soc. 2020. PMID: 32364710 Free PMC article.
References
-
- Krawczak M, Reiss J, Cooper DN. The mutational spectrum of single base-pair substitutions in mRNA splice junctions of human genes: causes and consequences. Hum. Genet. 1992;90:41–54. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

