Detection of miRNAs with a nanopore single-molecule counter
- PMID: 22845478
- PMCID: PMC3500609
- DOI: 10.1586/erm.12.58
Detection of miRNAs with a nanopore single-molecule counter
Abstract
miRNAs are short noncoding RNA molecules that are important in regulating gene expression. Due to the correlation of their expression levels and various diseases, miRNAs are being investigated as potential biomarkers for molecular diagnostics. The fast-growing miRNA exploration demands rapid, accurate, low-cost miRNA detection technologies. This article will focus on two platforms of nanopore single-molecule approach that can quantitatively measure miRNA levels in samples from tissue and cancer patient plasma. Both nanopore methods are sensitive and specific, and do not need labeling, enzymatic reaction or amplification. In the next 5 years, the nanopore-based miRNA techniques will be improved and validated for noninvasive and early diagnosis of diseases.
Figures
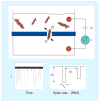


References
-
- Kim VN, Han J, Siomi MC. Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol. 2009;10:126–139. A comprehensive review of small RNAs including miRNAs, siRNAs and piRNAs in terms of biogenesis pathways and their regulations, along with their functions at both the genome and the transcriptome level. - PubMed
-
- Lee RC, Feinbaum RL, Ambros V. The C. elegans heterochronic gene LIN-4 encodes small RNAs with antisense complementarity to lin-14. Cell. 1993;75:843–854. The first publication that identified a small RNA encoded by the LIN-4 locus that was associated to the developmental timing of the nematode Caenorhabditis elegans by binding in the 3′ untranslated region (UTR) of LIN-14 mRNA and negatively modulating the protein Lin-14. - PubMed
-
- Wightman B, Ha I, Ruvkun G. Posttranscriptional regulation of the heterochronic gene LIN-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell. 1993;75:855–862. - PubMed
Website
-
- miRBase. http://microrna.sanger.ac.uk.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
