Filament formation of the FtsZ/tubulin-like protein TubZ from the Bacillus cereus pXO1 plasmid
- PMID: 22847006
- PMCID: PMC3442541
- DOI: 10.1074/jbc.M112.373803
Filament formation of the FtsZ/tubulin-like protein TubZ from the Bacillus cereus pXO1 plasmid
Abstract
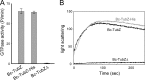
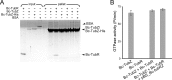
Stable maintenance of low-copy-number plasmids requires partition (par) systems that consist of a nucleotide hydrolase, a DNA-binding protein, and a cis-acting DNA-binding site. The FtsZ/tubulin-like GTPase TubZ was identified as a partitioning factor of the virulence plasmids pBtoxis and pXO1 in Bacillus thuringiensis and Bacillus anthracis, respectively. TubZ exhibits high GTPase activity and assembles into polymers both in vivo and in vitro, and its "treadmilling" movement is required for plasmid stability in the cell. To investigate the molecular mechanism of pXO1 plasmid segregation by TubZ filaments, we determined the crystal structures of Bacillus cereus TubZ in apo-, GDP-, and guanosine 5'-3-O-(thio)triphosphate (GTPγS)-bound forms at resolutions of 2.1, 1.9, and 3.3 Å, respectively. Interestingly, the slowly hydrolyzable GTP analog GTPγS was hydrolyzed to GDP in the crystal. In the post-GTP hydrolysis state, GDP-bound B. cereus TubZ forms a dimer by the head-to-tail association of individual subunits in the asymmetric unit, which is similar to the protofilament formation of FtsZ and B. thuringiensis TubZ. However, the M loop interacts with the nucleotide-binding site of the adjacent subunit and stabilizes the filament structure in a different manner, which indicates that the molecular assembly of the TubZ-related par systems is not stringently conserved. Furthermore, we show that the C-terminal tail of TubZ is required for association with the DNA-binding protein TubR. Using a combination of crystallography, site-directed mutagenesis, and biochemical analysis, our results provide the structural basis of the TubZ polymer that may drive DNA segregation.
Figures



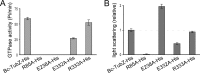
Similar articles
-
Cooperative DNA Binding of the Plasmid Partitioning Protein TubR from the Bacillus cereus pXO1 Plasmid.J Mol Biol. 2018 Dec 7;430(24):5015-5028. doi: 10.1016/j.jmb.2018.11.001. Epub 2018 Nov 8. J Mol Biol. 2018. PMID: 30414406
-
Crystallization and preliminary X-ray data analysis of the pXO1 plasmid-partitioning factor TubZ from Bacillus cereus.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012 Dec 1;68(Pt 12):1550-3. doi: 10.1107/S1744309112045551. Epub 2012 Nov 19. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2012. PMID: 23192045 Free PMC article.
-
Plasmid protein TubR uses a distinct mode of HTH-DNA binding and recruits the prokaryotic tubulin homolog TubZ to effect DNA partition.Proc Natl Acad Sci U S A. 2010 Jun 29;107(26):11763-8. doi: 10.1073/pnas.1003817107. Epub 2010 Jun 4. Proc Natl Acad Sci U S A. 2010. PMID: 20534443 Free PMC article.
-
Tubulin-Like Proteins in Prokaryotic DNA Positioning.Subcell Biochem. 2017;84:323-356. doi: 10.1007/978-3-319-53047-5_11. Subcell Biochem. 2017. PMID: 28500531 Review.
-
What sets Bacillus anthracis apart from other Bacillus species?Annu Rev Microbiol. 2009;63:451-76. doi: 10.1146/annurev.micro.091208.073255. Annu Rev Microbiol. 2009. PMID: 19514852 Review.
Cited by
-
Bacterial tubulin TubZ-Bt transitions between a two-stranded intermediate and a four-stranded filament upon GTP hydrolysis.Proc Natl Acad Sci U S A. 2014 Mar 4;111(9):3407-12. doi: 10.1073/pnas.1318339111. Epub 2014 Feb 18. Proc Natl Acad Sci U S A. 2014. PMID: 24550513 Free PMC article.
-
The C-terminal region of the plasmid partitioning protein TubY is a tetramer that can bind membranes and DNA.J Biol Chem. 2020 Dec 18;295(51):17770-17780. doi: 10.1074/jbc.RA120.014705. J Biol Chem. 2020. PMID: 33454013 Free PMC article.
-
Structure of the tubulin/FtsZ-like protein TubZ from Pseudomonas bacteriophage ΦKZ.J Mol Biol. 2013 Jun 26;425(12):2164-73. doi: 10.1016/j.jmb.2013.03.019. Epub 2013 Mar 22. J Mol Biol. 2013. PMID: 23528827 Free PMC article.
-
Catching a Walker in the Act-DNA Partitioning by ParA Family of Proteins.Front Microbiol. 2022 May 26;13:856547. doi: 10.3389/fmicb.2022.856547. eCollection 2022. Front Microbiol. 2022. PMID: 35694299 Free PMC article. Review.
-
A novel transcriptional activator, tubX, is required for the stability of Bacillus sphaericus mosquitocidal plasmid pBsph.J Bacteriol. 2014 Dec;196(24):4304-14. doi: 10.1128/JB.01855-14. Epub 2014 Sep 29. J Bacteriol. 2014. PMID: 25266379 Free PMC article.
References
-
- Carballido-López R., Errington J. (2003) A dynamic bacterial cytoskeleton. Trends Cell Biol. 13, 577–583 - PubMed
-
- Graumann P. L. (2007) Cytoskeletal elements in bacteria. Annu. Rev. Microbiol. 61, 589–618 - PubMed
-
- Cabeen M. T., Jacobs-Wagner C. (2010) The bacterial cytoskeleton. Annu. Rev. Genet. 44, 365–392 - PubMed
-
- van den Ent F., Amos L., Löwe J. (2001) Bacterial ancestry of actin and tubulin. Curr. Opin. Microbiol. 4, 634–638 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

