DNA of a circular minichromosome linearized by restriction enzymes or other reagents is resistant to further cleavage: an influence of chromatin topology on the accessibility of DNA
- PMID: 22848103
- PMCID: PMC3479189
- DOI: 10.1093/nar/gks723
DNA of a circular minichromosome linearized by restriction enzymes or other reagents is resistant to further cleavage: an influence of chromatin topology on the accessibility of DNA
Abstract
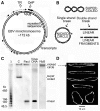
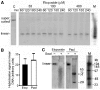
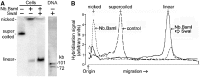
The accessibility of DNA in chromatin is an essential factor in regulating its activities. We studied the accessibility of the DNA in a ∼170 kb circular minichromosome to DNA-cleaving reagents using pulsed-field gel electrophoresis and fibre-fluorescence in situ hybridization on combed DNA molecules. Only one of several potential sites in the minichromosome DNA was accessible to restriction enzymes in permeabilized cells, and in growing cells only a single site at an essentially random position was cut by poisoned topoisomerase II, neocarzinostatin and γ-radiation, which have multiple potential cleavage sites; further sites were then inaccessible in the linearized minichromosomes. Sequential exposure to combinations of these reagents also resulted in cleavage at only a single site. Minichromosome DNA containing single-strand breaks created by a nicking endonuclease to relax any unconstrained superhelicity was also cut at only a single position by a restriction enzyme. Further sites became accessible after ≥95% of histones H2A, H2B and H1, and most non-histone proteins were extracted. These observations suggest that a global rearrangement of the three-dimensional packing and interactions of nucleosomes occurs when a circular minichromosome is linearized and results in its DNA becoming inaccessible to probes.
Figures





Similar articles
-
Repair of DNA strand breaks in a minichromosome in vivo: kinetics, modeling, and effects of inhibitors.PLoS One. 2013;8(1):e52966. doi: 10.1371/journal.pone.0052966. Epub 2013 Jan 30. PLoS One. 2013. PMID: 23382828 Free PMC article.
-
Structural changes in simian virus 40 chromatin as probed by restriction endonucleases.J Virol. 1979 Sep;31(3):718-32. doi: 10.1128/JVI.31.3.718-732.1979. J Virol. 1979. PMID: 229248 Free PMC article.
-
Structural organization of the hepatitis B virus minichromosome.J Mol Biol. 2001 Mar 16;307(1):183-96. doi: 10.1006/jmbi.2000.4481. J Mol Biol. 2001. PMID: 11243813
-
DNA repair in the context of chromatin.Cell Cycle. 2005 Apr;4(4):568-71. Epub 2005 Apr 21. Cell Cycle. 2005. PMID: 15753656 Review.
-
Nicking endonucleases.Biochemistry (Mosc). 2009 Dec;74(13):1457-66. doi: 10.1134/s0006297909130033. Biochemistry (Mosc). 2009. PMID: 20210703 Review.
Cited by
-
Intercalation of small molecules into DNA in chromatin is primarily controlled by superhelical constraint.PLoS One. 2019 Nov 20;14(11):e0224936. doi: 10.1371/journal.pone.0224936. eCollection 2019. PLoS One. 2019. PMID: 31747397 Free PMC article.
-
Chromatin remodeling modulates radiosensitivity of the daughter cells derived from cell population exposed to low- and high-LET irradiation.Oncotarget. 2017 Apr 20;8(32):52823-52836. doi: 10.18632/oncotarget.17275. eCollection 2017 Aug 8. Oncotarget. 2017. PMID: 28881774 Free PMC article.
-
Repair of DNA strand breaks in a minichromosome in vivo: kinetics, modeling, and effects of inhibitors.PLoS One. 2013;8(1):e52966. doi: 10.1371/journal.pone.0052966. Epub 2013 Jan 30. PLoS One. 2013. PMID: 23382828 Free PMC article.
-
Endogenous single-strand DNA breaks at RNA polymerase II promoters in Saccharomyces cerevisiae.Nucleic Acids Res. 2018 Nov 16;46(20):10649-10668. doi: 10.1093/nar/gky743. Nucleic Acids Res. 2018. PMID: 30445637 Free PMC article.
-
Nucleosome-positioning sequence repeats impact chromatin silencing in yeast minichromosomes.Genetics. 2014 Nov;198(3):1015-29. doi: 10.1534/genetics.114.169508. Epub 2014 Sep 3. Genetics. 2014. PMID: 25189873 Free PMC article.

