Comparative oncogenomics implicates the neurofibromin 1 gene (NF1) as a breast cancer driver
- PMID: 22851646
- PMCID: PMC3454871
- DOI: 10.1534/genetics.112.142802
Comparative oncogenomics implicates the neurofibromin 1 gene (NF1) as a breast cancer driver
Abstract
Identifying genomic alterations driving breast cancer is complicated by tumor diversity and genetic heterogeneity. Relevant mouse models are powerful for untangling this problem because such heterogeneity can be controlled. Inbred Chaos3 mice exhibit high levels of genomic instability leading to mammary tumors that have tumor gene expression profiles closely resembling mature human mammary luminal cell signatures. We genomically characterized mammary adenocarcinomas from these mice to identify cancer-causing genomic events that overlap common alterations in human breast cancer. Chaos3 tumors underwent recurrent copy number alterations (CNAs), particularly deletion of the RAS inhibitor Neurofibromin 1 (Nf1) in nearly all cases. These overlap with human CNAs including NF1, which is deleted or mutated in 27.7% of all breast carcinomas. Chaos3 mammary tumor cells exhibit RAS hyperactivation and increased sensitivity to RAS pathway inhibitors. These results indicate that spontaneous NF1 loss can drive breast cancer. This should be informative for treatment of the significant fraction of patients whose tumors bear NF1 mutations.
Figures
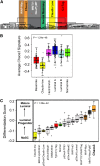

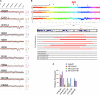
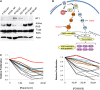

References
-
- Arima Y., Hayashi H., Kamata K., Goto T. M., Sasaki M., et al. , 2009. Decreased expression of neurofibromin contributes to epithelial-mesenchymal transition in neurofibromatosis type 1. Exp. Dermatol. 19: e136–e141 - PubMed
-
- Bajenaru M. L., Donahoe J., Corral T., Reilly K. M., Brophy S., et al. , 2001. Neurofibromatosis 1 (NF1) heterozygosity results in a cell-autonomous growth advantage for astrocytes. Glia 33: 314–323 - PubMed
-
- Carey L. A., Perou C. M., Livasy C. A., Dressler L. G., Cowan D., et al. , 2006. Race, breast cancer subtypes, and survival in the Carolina Breast Cancer Study. JAMA 295: 2492–2502 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

