Mixed exciton-charge-transfer states in photosystem II: Stark spectroscopy on site-directed mutants
- PMID: 22853895
- PMCID: PMC3400784
- DOI: 10.1016/j.bpj.2012.06.026
Mixed exciton-charge-transfer states in photosystem II: Stark spectroscopy on site-directed mutants
Abstract
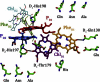
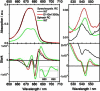
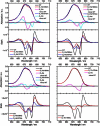
We investigated the electronic structure of the photosystem II reaction center (PSII RC) in relation to the light-induced charge separation process using Stark spectroscopy on a series of site-directed PSII RC mutants from the cyanobacterium Synechocystis sp. PCC 6803. The site-directed mutations modify the protein environment of the cofactors involved in charge separation (P(D1), P(D2), Chl(D1), and Phe(D1)). The results demonstrate that at least two different exciton states are mixed with charge-transfer (CT) states, yielding exciton states with CT character: (P(D2)(δ)(+)P(D1)(δ)(-)Chl(D1)) (673 nm) and (Chl(D1)(δ)(+)Phe(D1)(δ)(-)) (681 nm) (where the subscript indicates the wavelength of the electronic transition). Moreover, the CT state P(D2)(+)P(D1)(-) acquires excited-state character due to its mixing with an exciton state, producing (P(D2)(+)P(D1)(-))(δ) (684 nm). We conclude that the states that initiate charge separation are mixed exciton-CT states, and that the degree of mixing between exciton and CT states determines the efficiency of charge separation. In addition, the results reveal that the pigment-protein interactions fine-tune the energy of the exciton and CT states, and hence the mixing between these states. This mixing ultimately controls the selection and efficiency of a specific charge separation pathway, and highlights the capacity of the protein environment to control the functionality of the PSII RC complex.
Copyright © 2012 Biophysical Society. Published by Elsevier Inc. All rights reserved.
Figures
Similar articles
-
Site-directed mutations at D1-Thr179 of photosystem II in Synechocystis sp. PCC 6803 modify the spectroscopic properties of the accessory chlorophyll in the D1-branch of the reaction center.Biochemistry. 2008 Mar 11;47(10):3143-54. doi: 10.1021/bi702059f. Epub 2008 Feb 16. Biochemistry. 2008. PMID: 18278871
-
Spectroscopic properties of reaction center pigments in photosystem II core complexes: revision of the multimer model.Biophys J. 2008 Jul;95(1):105-19. doi: 10.1529/biophysj.107.123935. Epub 2008 Mar 13. Biophys J. 2008. PMID: 18339736 Free PMC article.
-
P680 (P(D1)P(D2)) and Chl(D1) as alternative electron donors in photosystem II core complexes and isolated reaction centers.J Photochem Photobiol B. 2011 Jul-Aug;104(1-2):44-50. doi: 10.1016/j.jphotobiol.2011.02.003. Epub 2011 Mar 5. J Photochem Photobiol B. 2011. PMID: 21377375
-
Site-directed mutations at D1-His198 and D2-His197 of photosystem II in Synechocystis PCC 6803: sites of primary charge separation and cation and triplet stabilization.Biochemistry. 2001 Aug 7;40(31):9265-81. doi: 10.1021/bi010121r. Biochemistry. 2001. PMID: 11478894
-
Primary radical ion pairs in photosystem II core complexes.Biochemistry (Mosc). 2014 Mar;79(3):197-204. doi: 10.1134/S0006297914030043. Biochemistry (Mosc). 2014. PMID: 24821445 Review.
Cited by
-
A TDDFT investigation of the Photosystem II reaction center: Insights into the precursors to charge separation.Proc Natl Acad Sci U S A. 2020 Aug 18;117(33):19705-19712. doi: 10.1073/pnas.1922158117. Epub 2020 Aug 3. Proc Natl Acad Sci U S A. 2020. PMID: 32747579 Free PMC article.
-
Quantum Coherence in Photosynthesis for Efficient Solar Energy Conversion.Nat Phys. 2014 Sep 1;10(9):676-682. doi: 10.1038/nphys3017. Nat Phys. 2014. PMID: 26870153 Free PMC article.
-
Femtosecond visible transient absorption spectroscopy of chlorophyll-f-containing photosystem II.Proc Natl Acad Sci U S A. 2020 Sep 15;117(37):23158-23164. doi: 10.1073/pnas.2006016117. Epub 2020 Aug 31. Proc Natl Acad Sci U S A. 2020. PMID: 32868421 Free PMC article.
-
A sixty-year tryst with photosynthesis and related processes: an informal personal perspective.Photosynth Res. 2019 Mar;139(1-3):15-43. doi: 10.1007/s11120-018-0590-0. Epub 2018 Oct 20. Photosynth Res. 2019. PMID: 30343396
-
Defining the far-red limit of photosystem I: the primary charge separation is functional to 840 nm.J Biol Chem. 2014 Aug 29;289(35):24630-9. doi: 10.1074/jbc.M114.555649. Epub 2014 Jul 14. J Biol Chem. 2014. PMID: 25023284 Free PMC article.
References
-
- Zouni A., Witt H.T., Orth P. Crystal structure of photosystem II from Synechococcus elongatus at 3.8 A resolution. Nature. 2001;409:739–743. - PubMed
-
- Ferreira K.N., Iverson T.M., Iwata S. Architecture of the photosynthetic oxygen-evolving center. Science. 2004;303:1831–1838. - PubMed
-
- Loll B., Kern J., Biesiadka J. Towards complete cofactor arrangement in the 3.0 A resolution structure of photosystem II. Nature. 2005;438:1040–1044. - PubMed
-
- Guskov A., Kern J., Saenger W. Cyanobacterial photosystem II at 2.9-A resolution and the role of quinones, lipids, channels and chloride. Nat. Struct. Mol. Biol. 2009;16:334–342. - PubMed
-
- Umena Y., Kawakami K., Kamiya N. Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature. 2011;473:55–60. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

