Multiple pathways of duplication formation with and without recombination (RecA) in Salmonella enterica
- PMID: 22865732
- PMCID: PMC3454872
- DOI: 10.1534/genetics.112.142570
Multiple pathways of duplication formation with and without recombination (RecA) in Salmonella enterica
Abstract
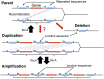
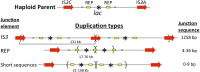
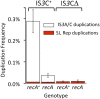
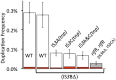
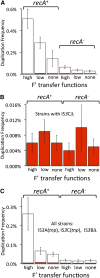
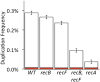
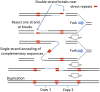
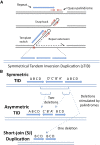
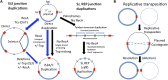
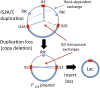
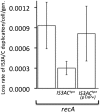
Duplications are often attributed to "unequal recombination" between separated, directly repeated sequence elements (>100 bp), events that leave a recombinant element at the duplication junction. However, in the bacterial chromosome, duplications form at high rates (10(-3)-10(-5)/cell/division) even without recombination (RecA). Here we describe 1800 spontaneous lac duplications trapped nonselectively on the low-copy F'(128) plasmid, where lac is flanked by direct repeats of the transposable element IS3 (1258 bp) and by numerous quasipalindromic REP elements (30 bp). Duplications form at a high rate (10(-4)/cell/division) that is reduced only about 11-fold in the absence of RecA. With and without RecA, most duplications arise by recombination between IS3 elements (97%). Formation of these duplications is stimulated by IS3 transposase (Tnp) and plasmid transfer functions (TraI). Three duplication pathways are proposed. First, plasmid dimers form at a high rate stimulated by RecA and are then modified by deletions between IS3 elements (resolution) that leave a monomeric plasmid with an IS3-flanked lac duplication. Second, without RecA, duplications occur by single-strand annealing of DNA ends generated in different sister chromosomes after transposase nicks DNA near participating IS3 elements. The absence of RecA may stimulate annealing by allowing chromosome breaks to persist. Third, a minority of lac duplications (3%) have short (0-36 bp) junction sequences (SJ), some of which are located within REP elements. These duplication types form without RecA, Tnp, or Tra by a pathway in which the palindromic junctions of a tandem inversion duplication (TID) may stimulate deletions that leave the final duplication.
Figures



Similar articles
-
Recombination and annealing pathways compete for substrates in making rrn duplications in Salmonella enterica.Genetics. 2014 Jan;196(1):119-35. doi: 10.1534/genetics.113.158519. Epub 2013 Nov 8. Genetics. 2014. PMID: 24214339 Free PMC article.
-
The effect of genomic position on reversion of a lac frameshift mutation (lacIZ33) during non-lethal selection (adaptive mutation).Mol Microbiol. 2002 May;44(4):1017-32. doi: 10.1046/j.1365-2958.2002.02934.x. Mol Microbiol. 2002. PMID: 12010495
-
The tandem inversion duplication in Salmonella enterica: selection drives unstable precursors to final mutation types.Genetics. 2010 May;185(1):65-80. doi: 10.1534/genetics.110.114074. Epub 2010 Mar 9. Genetics. 2010. PMID: 20215473 Free PMC article.
-
recA-independent DNA recombination between repetitive sequences: mechanisms and implications.Prog Nucleic Acid Res Mol Biol. 1996;54:253-92. doi: 10.1016/s0079-6603(08)60365-7. Prog Nucleic Acid Res Mol Biol. 1996. PMID: 8768077 Review. No abstract available.
-
Multiple pathways of deletion formation in Escherichia coli.Mutat Res. 1993 May;299(3-4):193-209. doi: 10.1016/0165-1218(93)90096-v. Mutat Res. 1993. PMID: 7683087 Review.
Cited by
-
Extensive amplification of GI-VII-6, a multidrug resistance genomic island of Salmonella enterica serovar Typhimurium, increases resistance to extended-spectrum cephalosporins.Front Microbiol. 2015 Feb 10;6:78. doi: 10.3389/fmicb.2015.00078. eCollection 2015. Front Microbiol. 2015. PMID: 25713569 Free PMC article.
-
Intraclonal Genome Stability of the Metallo-β-lactamase SPM-1-producing Pseudomonas aeruginosa ST277, an Endemic Clone Disseminated in Brazilian Hospitals.Front Microbiol. 2016 Dec 5;7:1946. doi: 10.3389/fmicb.2016.01946. eCollection 2016. Front Microbiol. 2016. PMID: 27994579 Free PMC article.
-
Rapid Evolution of Citrate Utilization by Escherichia coli by Direct Selection Requires citT and dctA.J Bacteriol. 2016 Feb 1;198(7):1022-34. doi: 10.1128/JB.00831-15. J Bacteriol. 2016. PMID: 26833416 Free PMC article.
-
Intrinsic adaptive value and early fate of gene duplication revealed by a bottom-up approach.Elife. 2018 Jan 5;7:e29739. doi: 10.7554/eLife.29739. Elife. 2018. PMID: 29303479 Free PMC article.
-
Three concurrent mechanisms generate gene copy number variation and transient antibiotic heteroresistance.Nat Commun. 2024 May 10;15(1):3981. doi: 10.1038/s41467-024-48233-0. Nat Commun. 2024. PMID: 38730266 Free PMC article.
References
-
- Anderson R. P., Roth J. R., 1978. Tandem chromosomal duplications in Salmonella typhimurium: fusion of histidine genes to novel promoters. J. Mol. Biol. 119: 147–166 - PubMed
-
- Andersson D. I., Slechta E. S., Roth J. R., 1998. Evidence that gene amplification underlies adaptive mutability of the bacterial lac operon. Science 282: 1133–1135 - PubMed
-
- Andersson D. I., Hughes D., Roth J. R. (), 2011. The Origin of Mutants Under Selection: Interactions of Mutation, Growth, and Selection. ASM Press, Washington, DC. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

