Molecular characterization of the glauce mutant: a central cell-specific function is required for double fertilization in Arabidopsis
- PMID: 22872756
- PMCID: PMC3462630
- DOI: 10.1105/tpc.112.096420
Molecular characterization of the glauce mutant: a central cell-specific function is required for double fertilization in Arabidopsis
Abstract
Double fertilization of the egg cell and the central cell by two sperm cells, resulting in the formation of the embryo and the endosperm, respectively, is a defining characteristic of flowering plants. The Arabidopsis thaliana female gametophytic mutant glauce (glc) can exhibit embryo development without any endosperm. Here, we show that in glc mutant embryo sacs one sperm cell successfully fuses with the egg cell but the second sperm cell fails to fuse with the central cell, resulting in single fertilization. Complementation studies using genes from the glc deletion interval identified an unusual genomic locus having homology to BAHD (for BEAT, AHCT, HCBT, and DAT) acyl-transferases with dual transcription units and alternative splicing that could rescue the sterility defect of glc. Expression of these transcripts appears restricted to the central cell, and expression within the central cell is sufficient to restore fertility. We conclude that the central cell actively promotes its own fertilization by the sperm cell through a signaling mechanism involving products of At1g65450. Successful fertilization of the egg cell is not blocked in the glc mutant, suggesting that evolution of double fertilization in flowering plants involved acquisition of specific functions by the central cell to enable its role as a second female gamete.
Figures
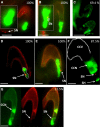






Similar articles
-
Arabidopsis GLAUCE promotes fertilization-independent endosperm development and expression of paternally inherited alleles.Development. 2007 Nov;134(22):4107-17. doi: 10.1242/dev.007310. Development. 2007. PMID: 17965055
-
The role of Arabidopsis thaliana NAR1, a cytosolic iron-sulfur cluster assembly component, in gametophytic gene expression and oxidative stress responses in vegetative tissue.New Phytol. 2013 Sep;199(4):925-935. doi: 10.1111/nph.12350. Epub 2013 Jun 5. New Phytol. 2013. PMID: 23734982
-
Expressing the diphtheria toxin A subunit from the HAP2(GCS1) promoter blocks sperm maturation and produces single sperm-like cells capable of fertilization.Plant Physiol. 2009 Nov;151(3):1390-400. doi: 10.1104/pp.109.144204. Epub 2009 Sep 4. Plant Physiol. 2009. PMID: 19734264 Free PMC article.
-
Gamete activation for fertilization and seed development in flowering plants.Curr Top Dev Biol. 2025;162:1-31. doi: 10.1016/bs.ctdb.2024.10.009. Epub 2024 Nov 13. Curr Top Dev Biol. 2025. PMID: 40180506 Review.
-
Epigenetic mechanisms governing seed development in plants.EMBO Rep. 2006 Dec;7(12):1223-7. doi: 10.1038/sj.embor.7400854. EMBO Rep. 2006. PMID: 17139298 Free PMC article. Review.
Cited by
-
Pollen tube entry into the synergid cell of Arabidopsis is observed at a site distinct from the filiform apparatus.Plant Reprod. 2013 Jun;26(2):93-9. doi: 10.1007/s00497-013-0211-1. Epub 2013 Feb 7. Plant Reprod. 2013. PMID: 23686222
-
The Armadillo repeat gene ZAK IXIK promotes Arabidopsis early embryo and endosperm development through a distinctive gametophytic maternal effect.Plant Cell. 2012 Oct;24(10):4026-43. doi: 10.1105/tpc.112.102384. Epub 2012 Oct 12. Plant Cell. 2012. PMID: 23064319 Free PMC article.
-
Genome-Wide Identification and Functional Characterization of the BAHD Acyltransferase Gene Family in Brassica napus L.Plants (Basel). 2025 Jul 15;14(14):2183. doi: 10.3390/plants14142183. Plants (Basel). 2025. PMID: 40733420 Free PMC article.
-
EGG CELL 1 contributes to egg-cell-dependent preferential fertilization in Arabidopsis.Nat Plants. 2024 Feb;10(2):268-282. doi: 10.1038/s41477-023-01616-5. Epub 2024 Jan 29. Nat Plants. 2024. PMID: 38287093
-
The cutin polymer matrix undergoes a fine architectural tuning from early tomato fruit development to ripening.Plant Physiol. 2022 Oct 27;190(3):1821-1840. doi: 10.1093/plphys/kiac392. Plant Physiol. 2022. PMID: 36018278 Free PMC article.
References
-
- Aw S.J., Hamamura Y., Chen Z., Schnittger A., Berger F. (2010). Sperm entry is sufficient to trigger division of the central cell but the paternal genome is required for endosperm development in Arabidopsis. Development 137: 2683–2690 - PubMed
-
- Baroux C., Spillane C., Grossniklaus U. (2002). Evolutionary origins of the endosperm in flowering plants. Genome Biol. 3: 1026.1–1026.5 - PubMed
-
- Berger F., Hamamura Y., Ingouff M., Higashiyama T. (2008). Double fertilization—Caught in the act. Trends Plant Sci. 13: 437–443 - PubMed
-
- Capron A., Gourgues M., Neiva L.S., Faure J.E., Berger F., Pagnussat G., Krishnan A., Alvarez-Mejia C., Vielle-Calzada J.P., Lee Y.R., Liu B., Sundaresan V. (2008). Maternal control of male-gamete delivery in Arabidopsis involves a putative GPI-anchored protein encoded by the LORELEI gene. Plant Cell 20: 3038–3049 - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

