RAPADILINO RECQL4 mutant protein lacks helicase and ATPase activity
- PMID: 22885111
- PMCID: PMC3500628
- DOI: 10.1016/j.bbadis.2012.07.014
RAPADILINO RECQL4 mutant protein lacks helicase and ATPase activity
Abstract
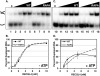
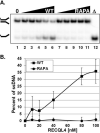
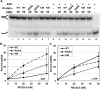
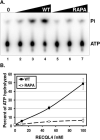
The RecQ family of helicases has been shown to play an important role in maintaining genomic stability. In humans, this family has five members and mutations in three of these helicases, BLM, WRN and RECQL4, are associated with disease. Alterations in RECQL4 are associated with three diseases, Rothmund-Thomson syndrome, Baller-Gerold syndrome, and RAPADILINO syndrome. One of the more common mutations found in RECQL4 is the RAPADILINO mutation, c.1390+2delT which is a splice-site mutation leading to an in-frame skipping of exon 7 resulting in 44 amino acids being deleted from the protein (p.Ala420-Ala463del). In order to characterize the RAPADILINO RECQL4 mutant protein, it was expressed in bacteria and purified using an established protocol. Strand annealing, helicase, and ATPase assays were conducted to characterize the protein's activities relative to WT RECQL4. Here we show that strand annealing activity in the absence of ATP is unchanged from that of WT RECQL4. However, the RAPADILINO protein variant lacks helicase and ssDNA-stimulated ATPase activity. These observations help explain the underlying molecular etiology of the disease and our findings provide insight into the genotype and phenotype association among RECQL4 syndromes.
Copyright © 2012. Published by Elsevier B.V.
Figures


Similar articles
-
Human RecQL4 helicase plays multifaceted roles in the genomic stability of normal and cancer cells.Cancer Lett. 2018 Jan 28;413:1-10. doi: 10.1016/j.canlet.2017.10.021. Epub 2017 Nov 7. Cancer Lett. 2018. PMID: 29080750 Review.
-
RECQL4 Regulates p53 Function In Vivo During Skeletogenesis.J Bone Miner Res. 2015 Jun;30(6):1077-89. doi: 10.1002/jbmr.2436. J Bone Miner Res. 2015. PMID: 25556649
-
The versatile RECQL4.Genet Med. 2006 Apr;8(4):213-6. doi: 10.1097/01.gim.0000214457.58378.1a. Genet Med. 2006. PMID: 16617241 Review.
-
The molecular role of the Rothmund-Thomson-, RAPADILINO- and Baller-Gerold-gene product, RECQL4: recent progress.Cell Mol Life Sci. 2007 Apr;64(7-8):796-802. doi: 10.1007/s00018-007-6468-5. Cell Mol Life Sci. 2007. PMID: 17364146 Free PMC article. Review.
-
Long-term follow-up and molecular characterization of a patient with a RECQL4 mutation spectrum disorder.Dermatology. 2013;226(4):353-7. doi: 10.1159/000351311. Epub 2013 Jul 26. Dermatology. 2013. PMID: 23899764
Cited by
-
RECQL4 requires PARP1 for recruitment to DNA damage, and PARG dePARylation facilitates its associated role in end joining.Exp Mol Med. 2025 Feb;57(1):264-280. doi: 10.1038/s12276-024-01383-z. Epub 2025 Jan 28. Exp Mol Med. 2025. PMID: 39870799 Free PMC article.
-
Congenital Diseases of DNA Replication: Clinical Phenotypes and Molecular Mechanisms.Int J Mol Sci. 2021 Jan 18;22(2):911. doi: 10.3390/ijms22020911. Int J Mol Sci. 2021. PMID: 33477564 Free PMC article. Review.
-
Molecular Mechanisms of the RECQ4 Pathogenic Mutations.Front Mol Biosci. 2021 Nov 18;8:791194. doi: 10.3389/fmolb.2021.791194. eCollection 2021. Front Mol Biosci. 2021. PMID: 34869606 Free PMC article. Review.
-
Human RecQ helicases in DNA repair, recombination, and replication.Annu Rev Biochem. 2014;83:519-52. doi: 10.1146/annurev-biochem-060713-035428. Epub 2014 Mar 3. Annu Rev Biochem. 2014. PMID: 24606147 Free PMC article. Review.
-
Immunodeficiency in a Child with Rapadilino Syndrome: A Case Report and Review of the Literature.Case Reports Immunol. 2015;2015:137368. doi: 10.1155/2015/137368. Epub 2015 May 6. Case Reports Immunol. 2015. PMID: 26064716 Free PMC article.
References
-
- Vindigni A, Marino F, Gileadi O. Probing the structural basis of RecQ helicase function. Biophys. Chem. 2010;149:67–77. - PubMed
-
- Wang LL, Gannavarapu A, Kozinetz CA, Levy ML, Lewis RA, Chintagumpala MM, Ruiz-Maldanado R, Contreras-Ruiz J, Cunniff C, Erickson RP, Lev D, Rogers M, Zackai EH, Plon SE. Association between osteosarcoma and deleterious mutations in the RECQL4 gene in Rothmund-Thomson syndrome. J. Natl. Cancer Inst. 2003;95:669–674. - PubMed
-
- Siitonen HA, Sotkasiira J, Biervliet M, Benmansour A, Capri Y, Cormier-Daire V, Crandall B, Hannula-Jouppi K, Hennekam R, Herzog D, Keymolen K, Lipsanen-Nyman M, Miny P, Plon SE, Riedl S, Sarkar A, Vargas FR, Verloes A, Wang LL, Kaariainen H, Kestila M. The mutation spectrum in RECQL4 diseases. Eur. J. Hum. Genet. 2009;17:151–158. - PMC - PubMed
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

