Avogadro: an advanced semantic chemical editor, visualization, and analysis platform
- PMID: 22889332
- PMCID: PMC3542060
- DOI: 10.1186/1758-2946-4-17
Avogadro: an advanced semantic chemical editor, visualization, and analysis platform
Abstract
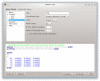
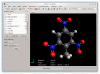
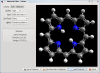
Background: The Avogadro project has developed an advanced molecule editor and visualizer designed for cross-platform use in computational chemistry, molecular modeling, bioinformatics, materials science, and related areas. It offers flexible, high quality rendering, and a powerful plugin architecture. Typical uses include building molecular structures, formatting input files, and analyzing output of a wide variety of computational chemistry packages. By using the CML file format as its native document type, Avogadro seeks to enhance the semantic accessibility of chemical data types.
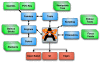
Results: The work presented here details the Avogadro library, which is a framework providing a code library and application programming interface (API) with three-dimensional visualization capabilities; and has direct applications to research and education in the fields of chemistry, physics, materials science, and biology. The Avogadro application provides a rich graphical interface using dynamically loaded plugins through the library itself. The application and library can each be extended by implementing a plugin module in C++ or Python to explore different visualization techniques, build/manipulate molecular structures, and interact with other programs. We describe some example extensions, one which uses a genetic algorithm to find stable crystal structures, and one which interfaces with the PackMol program to create packed, solvated structures for molecular dynamics simulations. The 1.0 release series of Avogadro is the main focus of the results discussed here.
Conclusions: Avogadro offers a semantic chemical builder and platform for visualization and analysis. For users, it offers an easy-to-use builder, integrated support for downloading from common databases such as PubChem and the Protein Data Bank, extracting chemical data from a wide variety of formats, including computational chemistry output, and native, semantic support for the CML file format. For developers, it can be easily extended via a powerful plugin mechanism to support new features in organic chemistry, inorganic complexes, drug design, materials, biomolecules, and simulations. Avogadro is freely available under an open-source license from http://avogadro.openmolecules.net.
Figures




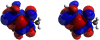

Similar articles
-
From data to analysis: linking NWChem and Avogadro with the syntax and semantics of Chemical Markup Language.J Cheminform. 2013 May 24;5(1):25. doi: 10.1186/1758-2946-5-25. J Cheminform. 2013. PMID: 23705910 Free PMC article.
-
Bioclipse: an open source workbench for chemo- and bioinformatics.BMC Bioinformatics. 2007 Feb 22;8:59. doi: 10.1186/1471-2105-8-59. BMC Bioinformatics. 2007. PMID: 17316423 Free PMC article.
-
Bioclipse 2: a scriptable integration platform for the life sciences.BMC Bioinformatics. 2009 Dec 3;10:397. doi: 10.1186/1471-2105-10-397. BMC Bioinformatics. 2009. PMID: 19958528 Free PMC article.
-
A tool written in Scala for preparation and analysis in MD simulation and 3D-RISM calculation of biomolecules.Biophys Physicobiol. 2019 Nov 29;16:485-489. doi: 10.2142/biophysico.16.0_485. eCollection 2019. Biophys Physicobiol. 2019. PMID: 31984200 Free PMC article. Review.
-
Array programming with NumPy.Nature. 2020 Sep;585(7825):357-362. doi: 10.1038/s41586-020-2649-2. Epub 2020 Sep 16. Nature. 2020. PMID: 32939066 Free PMC article. Review.
Cited by
-
Resolving the Size and Charge of Small Particles: A Predictive Model of Nanopore Mechanics.J Phys Chem C Nanomater Interfaces. 2024 Aug 9;128(41):17619-17630. doi: 10.1021/acs.jpcc.4c02722. eCollection 2024 Oct 17. J Phys Chem C Nanomater Interfaces. 2024. PMID: 39439880 Free PMC article.
-
Computational exploration of acefylline derivatives as MAO-B inhibitors for Parkinson's disease: insights from molecular docking, DFT, ADMET, and molecular dynamics approaches.Front Chem. 2024 Oct 8;12:1449165. doi: 10.3389/fchem.2024.1449165. eCollection 2024. Front Chem. 2024. PMID: 39439933 Free PMC article.
-
Unearthing the Real-Time Excited State Dynamics from Antenna to Rare Earth Ions Using Ultrafast Transient Absorption.JACS Au. 2024 Aug 20;4(10):3813-3822. doi: 10.1021/jacsau.4c00468. eCollection 2024 Oct 28. JACS Au. 2024. PMID: 39483220 Free PMC article.
-
Synthesis, Enzymatic Degradation, and Polymer-Miscibility Evaluation of Nonionic Antimicrobial Hyperbranched Polyesters with Indole or Isatin Functionalities.Biomacromolecules. 2021 May 10;22(5):2256-2271. doi: 10.1021/acs.biomac.1c00343. Epub 2021 Apr 26. Biomacromolecules. 2021. PMID: 33900740 Free PMC article.
-
Polymerization of silanes through dehydrogenative Si-Si bond formation on metal surfaces.Nat Chem. 2021 Apr;13(4):350-357. doi: 10.1038/s41557-021-00651-z. Epub 2021 Mar 29. Nat Chem. 2021. PMID: 33782562
References
-
- Hanson RM, Howard MT, Willighagen EL, Jmol: an open-source Java viewer for chemical structures in 3D. 2012. [ http://www.jmol.org]
-
- DeLano WL. The PyMOL Molecular Graphics System. 2002. [ http://www.pymol.org]
-
- Tarini M, Cignoni P, Montani C. Ambient Occlusion and Edge Cueing for Enhancing Real Time Molecular Visualization. IEEE Trans Visualization and Comput Graphics. 2006;12(5):1237–1244. - PubMed
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

