Pressure-assisted capillary electrophoresis coupling with matrix-assisted laser desorption/ionization-mass spectrometric imaging for quantitative analysis of complex peptide mixtures
- PMID: 22891936
- PMCID: PMC3445710
- DOI: 10.1021/ac300628s
Pressure-assisted capillary electrophoresis coupling with matrix-assisted laser desorption/ionization-mass spectrometric imaging for quantitative analysis of complex peptide mixtures
Abstract
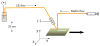
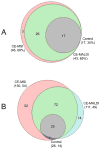
Herein, we report a pressure-assisted capillary electrophoresis-mass spectrometric imaging (PACE-MSI) platform for peptide analysis. This new platform has addressed the sample diffusion and peak splitting problems that appeared in our previous groove design, and it enables homogeneous deposition of the CE trace for high-throughput MALDI imaging. In the coupling of CE to MSI, individual peaks (m/z) can be visualized as discrete colored image regions and extracted from the MS imaging data, thus eliminating issues with peak overlapping and reducing reliance on an ultrahigh mass resolution mass spectrometer. Through a PACE separation, 46 tryptic peptides from bovine serum albumin and 150 putative neuropeptides from the pericardial organs of a model organism blue crab Callinectes sapidus were detected from the MALDI MS imaging traces, enabling a 4- to 6-fold increase of peptide coverage as compared with direct MALDI MS analysis. For the first time, quantitation with high accuracy was obtained using PACE-MSI for both digested tryptic peptides and endogenous neuropeptides from complex biological samples in combination with isotopic formaldehyde labeling. Although MSI is typically employed in tissue imaging, we show in this report that it offers a unique tool for quantitative analysis of complex trace-level analytes with CE separation. These results demonstrate a great potential of the PACE-MSI platform for enhanced quantitative proteomics and neuropeptidomics.
Figures



References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

